DesignSafe Use Cases
DesignSafe provides a wide variety of resources that allow researchers to effectively share, find, analyze, and publish data; perform numerical simulations and utilize high performance computing (HPC); and integrate diverse datasets.
To help users fully embrace DesignSafe functionalities, we have developed a suite of Use Cases that demonstrate how DesignSafe is being used to advance natural hazards research. Practical products, examples, and scripts developed as part of these Use Cases are provided at the links below. The different simulation codes, tools, and DesignSafe resources used in each Use Case are also indicated.
Data Analytics
- Damage Tagging of Field Images (Taggit, HazMapper)
- Machine Learning and AI Resources (Jupyter, Interactive Data Analytics, HPC)
- Visualization of spatially distributed data for risk and resilience analysis (Jupyter, Interactive Data Visualization)
Taggit - Image Tagging
Taggit User Guide: Basic Image Browsing and Mapping
Using HazMapper and Taggit to browse thumbnails of large numbers of images and map their locations
Fred Haan – Calvin University Key Words: Taggit, HazMapper, image browsing
Resources
The example makes use of the following DesignSafe resources:
Description
This user guide demonstrates how to use HazMapper and Taggit applications on DesignSafe to browse through large numbers of image files. HazMapper and Taggit should be considered different ways of viewing the same set of images. You see a thumbnail Gallery of those images when you use Taggit, and you see a Map of those images when you use HazMapper, but it is the same *.hazmapper file in both cases.
NOTE: You always start a Map/Gallery file in HazMapper. All browsing/mapping starts with HazMapper, and you can switch back and forth between the apps as you wish. Also, at this time you cannot publish a Map/Gallery but this functionality is coming soon.
Creating a Map/Gallery and Browsing Images
Taggit and HazMapper allow you to browse and map image files that are on DesignSafe. This document shows you how to get started with Taggit and HazMapper for a set of images available on DesignSafe.
As an example workflow, we will consider a damage survey dataset from the December 2021 Midwest tornado outbreak (see below).

If you were looking at this project on DesignSafe, you might be interested in seeing the image files in the folder shown below.
In order to browse thumbnail images of these photographs or see them laid out on a map, you can launch HazMapper to get started.
NOTE: All browsing/mapping starts with HazMapper. HazMapper and Taggit are just different ways of viewing the same Map/Gallery.
So to get started, we launch HazMapper from the Visualization tab of the Tools and Applications page of the DesignSafe Workspace.
Once HazMapper is up, click on “Create a New Map” as shown below:

Fill out a name and description of the “Map” you will create. This Map will also be a Gallery in Taggit. You then select a Save Location for your Map/Gallery by pulling down the menu on the right:
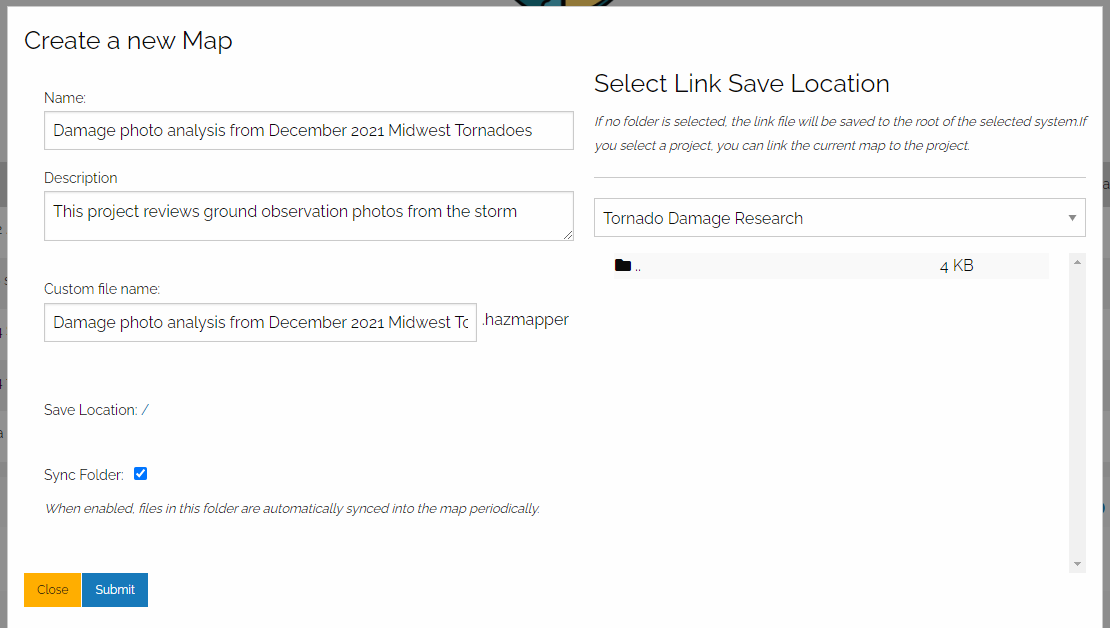
When you are selecting a Save Location, you should probably select from MyProjects. That way, you can share your Map/Gallery with other users. I selected a project called “Tornado Damage Research” as shown below.
Next you click on Assets and Import from DesignSafe to load the images you want to see:
I'm going to select files from the StEER – 10 December 2021 Midwest Tornado Outbreak project:
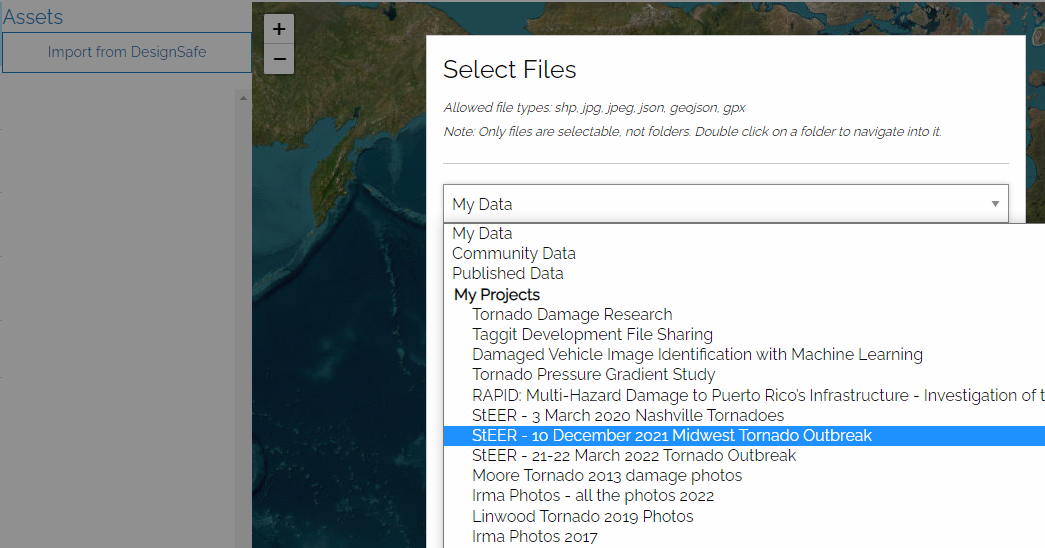
Once you navigate to the folder containing the image files you want, you can use shift-select to select multiple files to import:
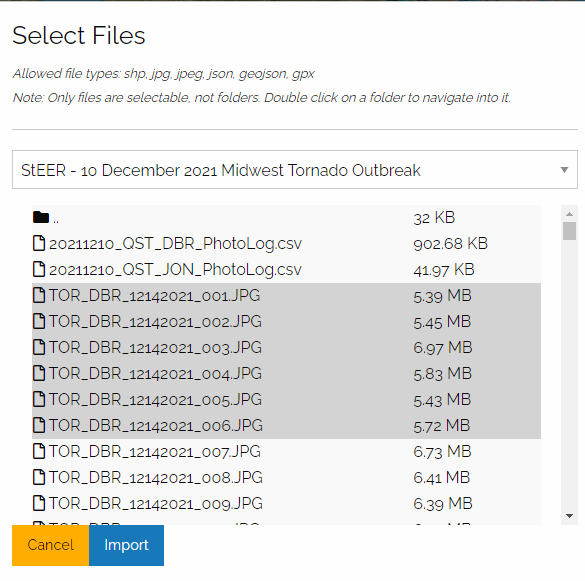
Once you press Import, you will see messages about successful imports on the right side of the screen:
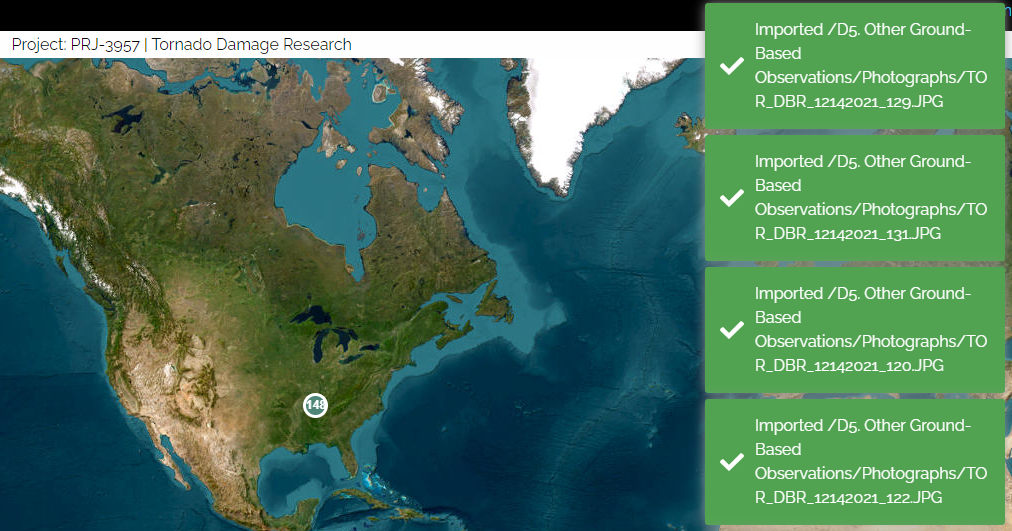
Browsing Thumbnail Images in Taggit
Once you've loaded your images, you can zoom in on your map to see the location of each photo. However, for this Taggit workflow demonstration, we will click on Manage and select View in Taggit as show below:
This will launch Taggit and show thumbnail images of all the photos that were just loaded in HazMapper. You can now browse through all the photos as shown below.
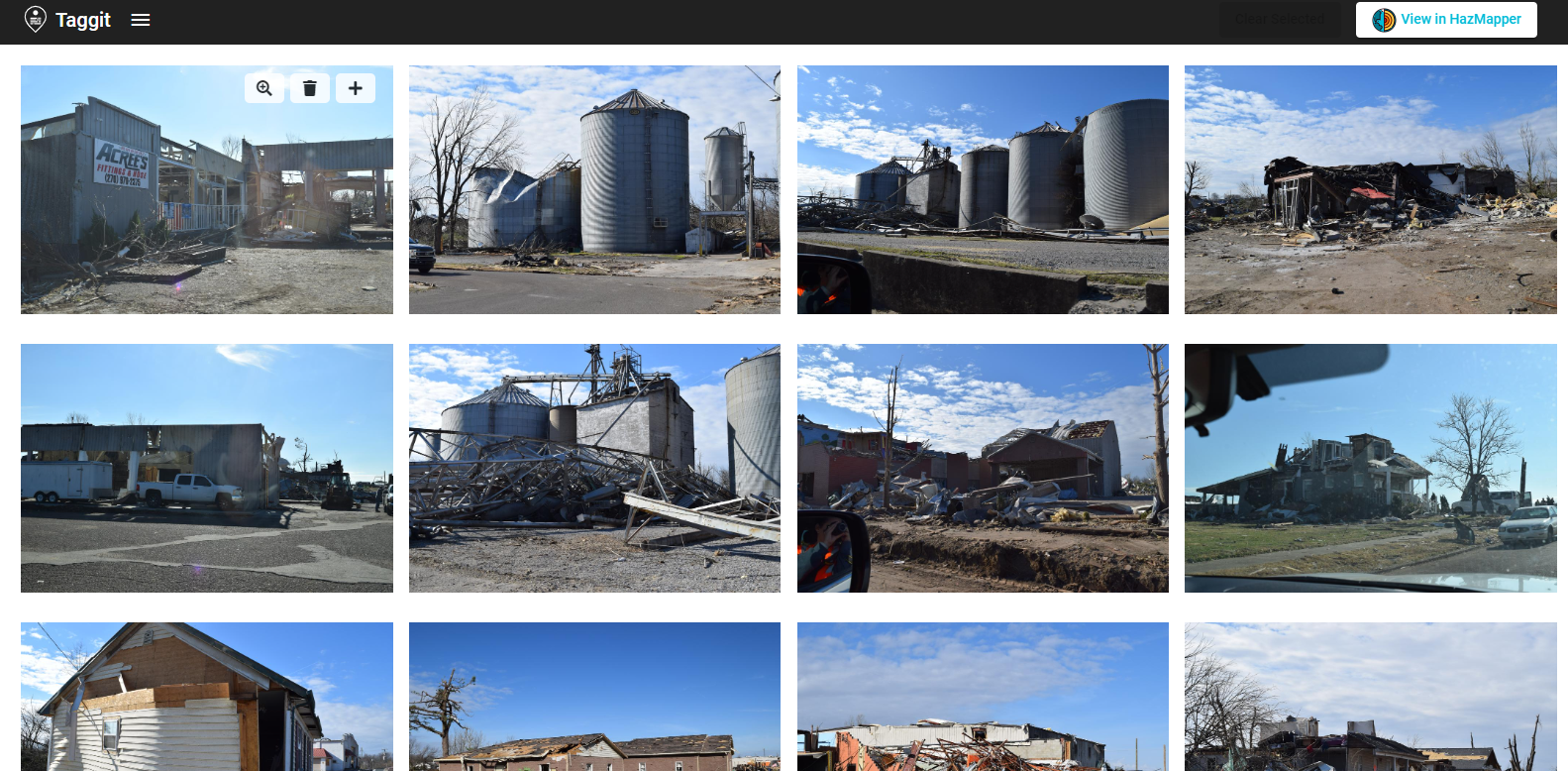

You can then add more photos to the Gallery using the pull-down menu in the upper-left corner. Selecting Import image from DesignSafe gives you access to all the files on DesignSafe.
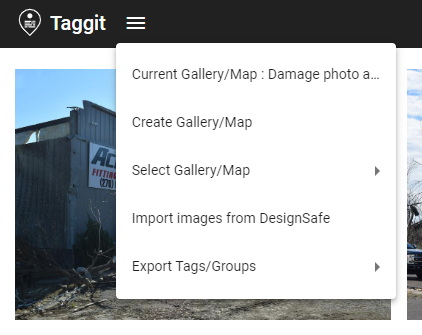
If you want to go back to HazMapper and put all the photos on a map, click the View in HazMapper button in the upper right corner of the screen.

References
- Kijewski-Correa et al. (2021) for PRJ-3349 StEER - 10 December 2021 Midwest Tornado Outbreak.
ML and AI
ML and AI
An Example-Based Introduction to Common Machine Learning Approaches
Joseph P. Vantassel and Wenyang Zhang, Texas Advanced Computing Center - The University of Texas at Austin
With the increasing acquisition and sharing of data in the natural hazards community, solutions from data science, in particular machine learning, are increasingly being applied to natural hazard problems. To better equip the natural hazards community to understand and utilize these solution this use case presents an example-based introduction to common machine learning approaches. This use case is not intended to be exhaustive in its coverage of machine learning approaches (as there are many), nor in its coverage of the selected approaches (as they are more complex than can be effectively communicated here), rather, this use case is intended to provide a high-level overview of different approaches to using machine learning to solve data-related problems. The example makes use of the following DesignSafe resources:
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
DesignSafe Resources
The following DesignSafe resources are leveraged in this example:
Geospatial data analysis and Visualization on DS - QGIS
Jupyter notebooks on DS Jupyterhub
Citation and Licensing
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
Please cite Durante and Rathje (2021) to acknowledge the use of any resources for the Random Forest and Neural Networks examples included in this use case.
-
This software is distributed under the GNU General Public License.
Overview of ML examples
This use case is example-based meaning that is its contents have been organized into self-contained examples. These self-contained example are organized by machine learning algorithm. Importantly, the machine learning algorithm applied to the specific example provided here are not the only (or even necessarily the optimal) algorithm for that particular (or related) problem, instead the datasets considered are used merely for example and the algorithm applied is but one of the potentially many reasonable alternatives one could use to solve that particular problem. The focus of these examples is to demonstrate the general procedure for applying that particular machine learning algorithm and does not necessarily indicate that this is the correct or optimal solution.
To run the examples for yourself, first copy the directory for the example you are interested in. You can
do this by following the links below to find the location of the associated notebooks in community data,
selecting the directory of interest (e.g., 0_linear_regression for the linear regression example) you will
need to navigate up one directory to make this selection and then selecting Copy > My Data > Copy Here. You
can then navigate to your My Data and run, explore, and modify the notebooks from your user space. If you do
not make a copy the notebooks will open as read-only and you will not be able to fully explore the examples provided.
Linear Regression
Linear regression seeks to find linear relationships between features in a dataset and an associated set of labels (i.e., real values to be predicted). Linear regression is one of the simplest machine learning algorithms and likely one that many natural hazards researchers will already be familiar with from undergraduate mathematics coursework (e.g., statistics, linear algebra). The example for linear regression presented in this use case shows the process of attempting to predict housing prices from house and neighborhood characteristics. The notebooks cover how to perform basic linear regression using the raw features, combine those features (also called feature crosses) to produce better predictions, use regularization to reduce overfitting, and use learning curves as a diagnostic tool for machine learning problems.
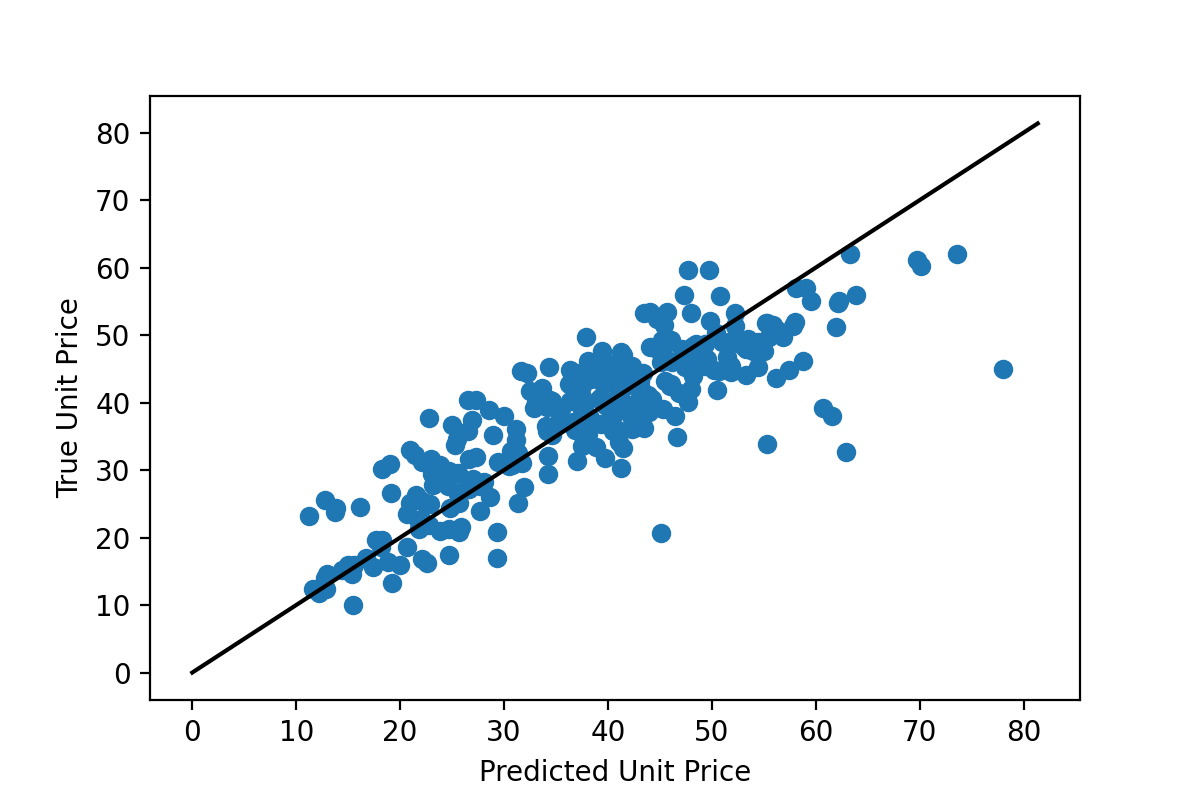
Open Jupyter Notebook dirctly:
- Linear Regression, Real Estate Data Set:
- Linear Regression, Real Estate Data Set - Learning Curves:
- Linear Regression, Real Estate Data Set - Regularization:
or View in the Data Depot: Linear Regression Example
Random Forest
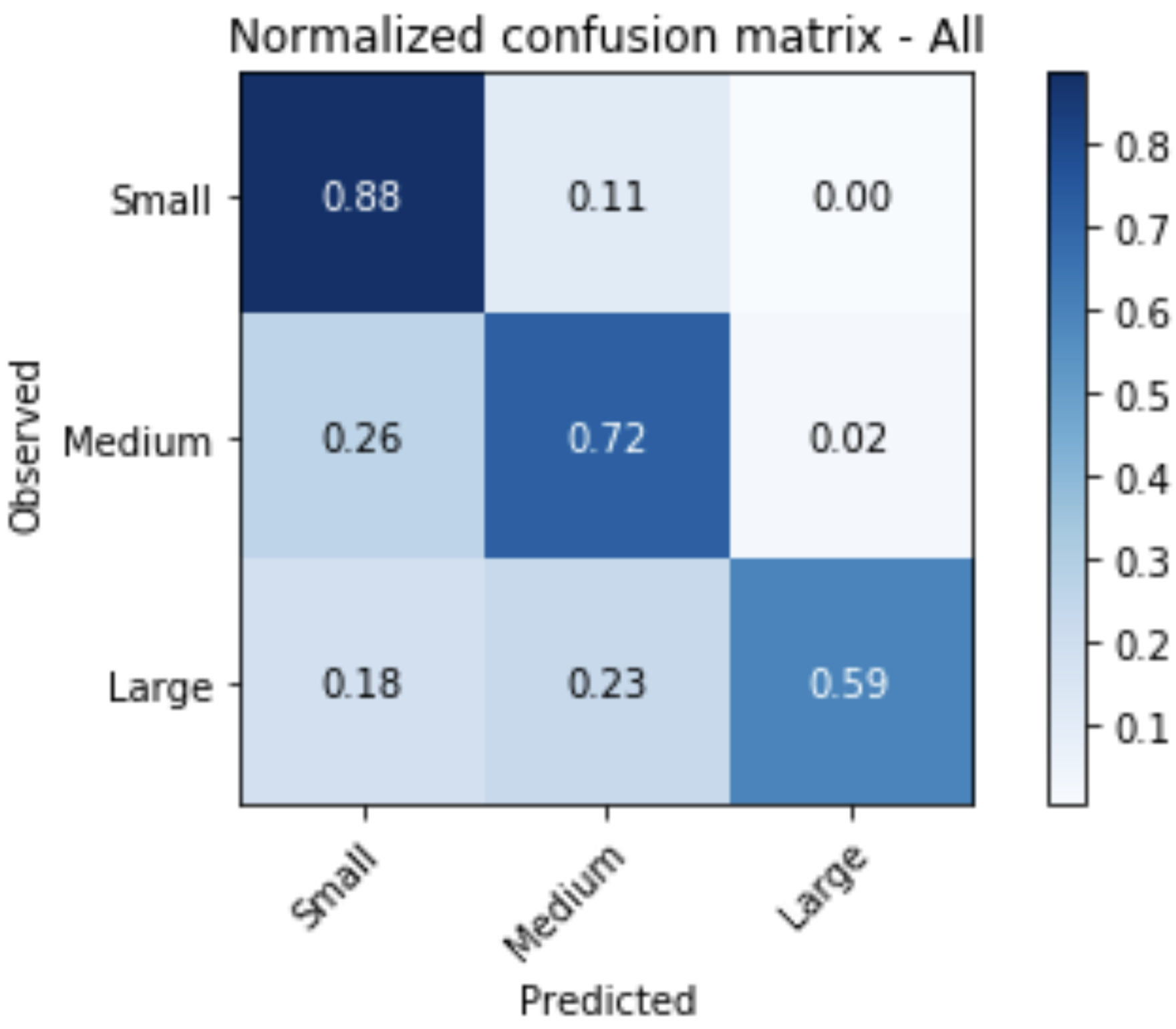
Random forests or random decision forests is an ensemble learning method for classification, regression and other tasks that operates by constructing a multitude of decision trees at training time. For classification tasks, the output of the random forest is the class selected by most trees. For regression tasks, the mean or average prediction of the individual trees is returned. Random decision forests correct for decision trees' habit of overfitting to their training set. Random forests generally outperform decision trees, but their accuracy is lower than gradient boosted trees. However, data characteristics can affect their performance.
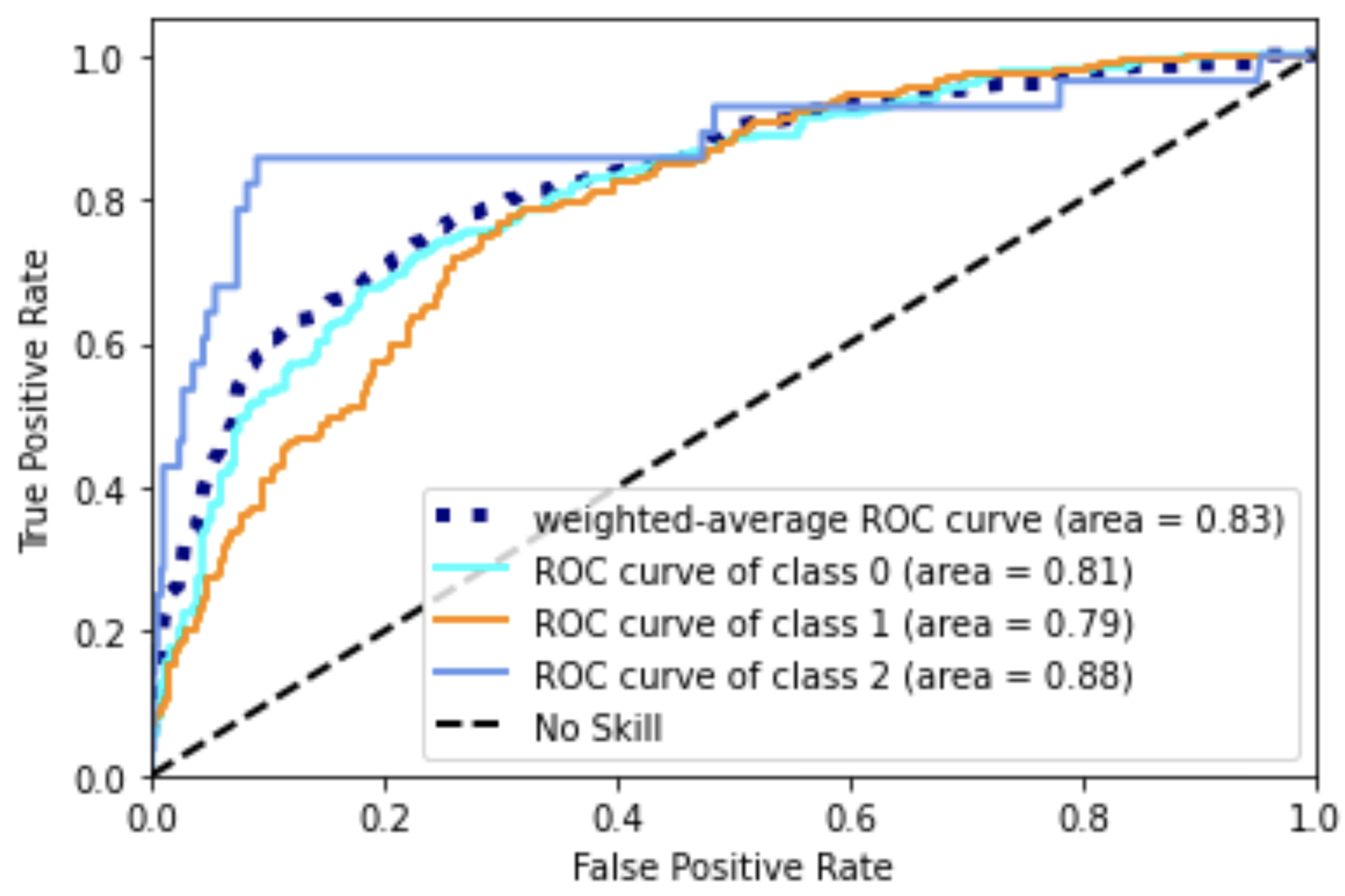
Open Jupyter Notebook directly:
or View in the Data Depot: Random Forest Example
Neural Networks
Artificial neural networks (ANNs), usually simply called neural networks (NNs), are computing systems inspired by the biological neural networks that constitute animal brains. An ANN is based on a collection of connected units or nodes called artificial neurons, which loosely model the neurons in a biological brain. Each connection, like the synapses in a biological brain, can transmit a signal to other neurons. An artificial neuron receives a signal then processes it and can signal neurons connected to it. The "signal" at a connection is a real number, and the output of each neuron is computed by some non-linear function of the sum of its inputs. The connections are called edges. Neurons and edges typically have a weight that adjusts as learning proceeds. The weight increases or decreases the strength of the signal at a connection. Neurons may have a threshold such that a signal is sent only if the aggregate signal crosses that threshold. Typically, neurons are aggregated into layers. Different layers may perform different transformations on their inputs. Signals travel from the first layer (the input layer), to the last layer (the output layer), possibly after traversing the layers multiple times.
Open Jupyter Notebook dirctly:
or View in the Data Depot: Artificial Neural Network Example
Convolutional Neural Networks
Convolutional neural networks fall under the deep learning subset of machine learning and are an effective tool for processing and understanding image and image-like data. The convolutional neural network example will show an image classification algorithm for automatically reading hand-written digits. The network will be provided an image of a hand-written digit and predict a label classifying it as a number between 0 and 9. The notebooks will show how to install Keras/TensorFlow, load a standard dataset, pre-process the data for acceptance by the network, design and train a convolutional neural network using Keras/TensorFlow, and visualize correct and incorrect output predictions. For those who have access to graphical processing unit (GPU) computational resources a replica of the main notebook is provided that can run across multiple GPUs on a single machine.

Open Jupyter Notebook dirctly:
- Convolutional Neural Networks, MNIST Dataset:
- Convolutional Neural Networks, MNIST Dataset - Parallel:
or View in the Data Depot: Convolutional Neural Network Example
Visualization of spatially distributed data
Visualization of Spatially Distributed Data
Jupyter notebook for visualization of spatially distributed data in risk and resilience analysis
Raul Rincon - Dept. of Civil and Environmental Engineering, Rice University
Jamie E. Padgett - Dept. of Civil and Environmental Engineering, Rice University
Keywords: visualization; risk and resilience; infrastructure systems; static, interactive, and animated maps and figures, effective communication
Resources
Jupyter Notebooks
The following Jupyter notebook is the basis for the use case described in this section. You can access and run it directly on DesignSafe by clicking on the "Open in DesignSafe" button.
| Scope | Notebook |
|---|---|
| visualization of spatially-distributed Data |
visualization_risk_resilience.ipynb |
DesignSafe Resources
The following DesignSafe resources were used in developing this use case.
Background
Citation and Licensing
-
Please cite Rincon and Padgett (2023) to acknowledge the use of resources from this use case.
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License.
Description
Effective visualization tools for communication of risk and resilience metrics are needed to translate technical information into trustable and useful outputs for decision-making, and also to aid in the research process, including sanity checks, verification, and validation steps, among others. Considering the diverse purposes, it may be desired to explore input, intermediate results, or final outcomes during the risk and resilience assessment (see Figure 1). This use case addresses these needs by leveraging different Python libraries to visualize spatially distributed data, especially focusing on risk and resilience analysis products. Some of the visualized products represent data that can be obtained through the Tools & Applications space in the DesignSafe cyberinfrastructure (for example using the SimCenter research tools) or other platforms for resilience measurement science (see IN-CORE platform). Procedures to develop static, interactive, and animated figures and maps are presented throughout the use case. The created maps aim to visualize hazard-to-resilience outputs obtained from the analysis of regionally distributed systems. Hence this Jupyter notebook may serve as a launching point for other researchers to adapt code for visualizing various stages along the regional risk and resilience quantification workflow.
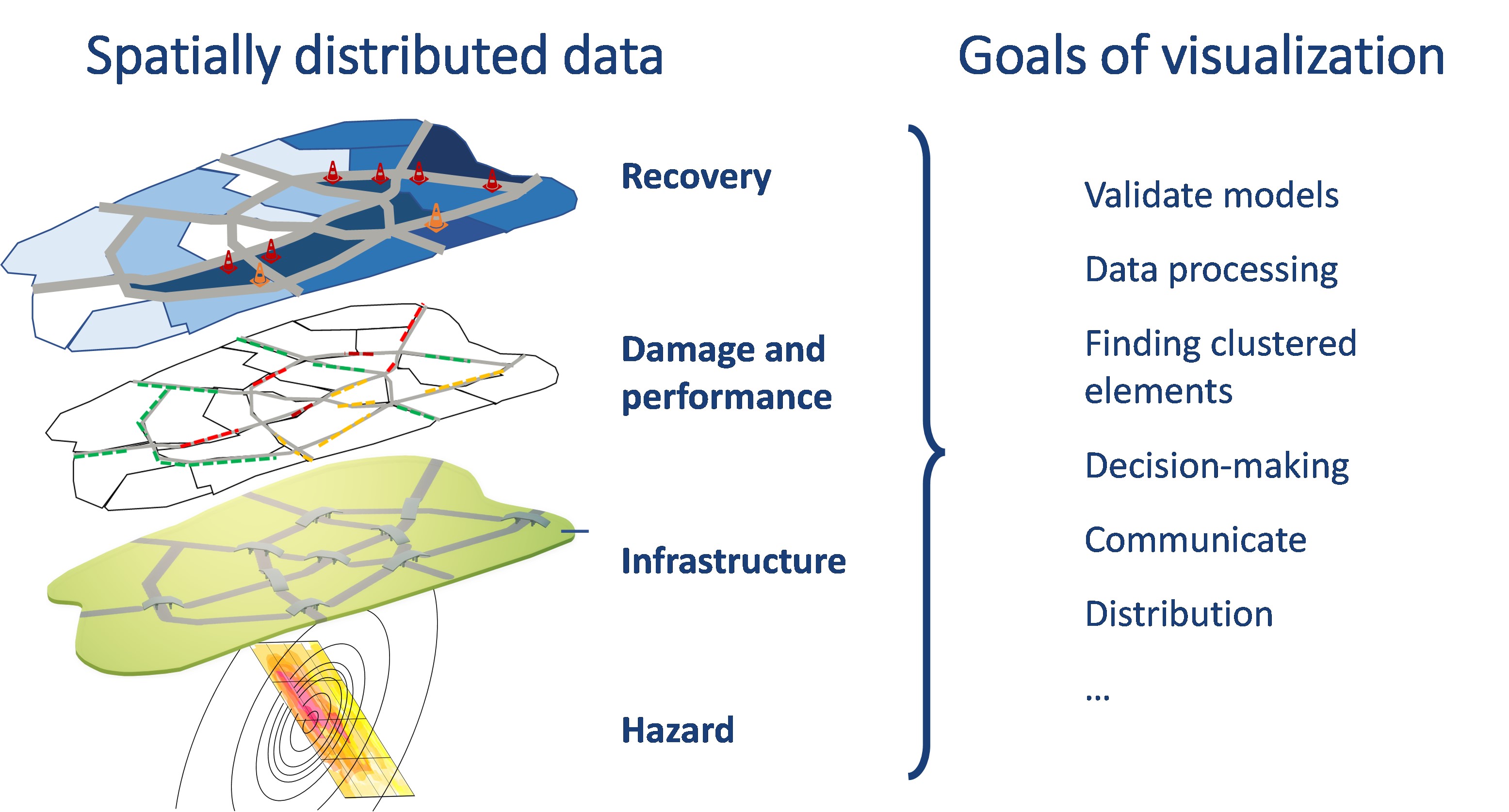
Figure 1. Risk and resilience outputs and stakeholder visualization needs
Implementation
This use case adopts a representative hazard and distributed infrastructure system for illustration purposes. A sparse representation of the highway network in the Memphis Metropolitan and Statistical Area (MMSA) subjected to a point-source earthquake scenario is considered. The highway network is defined using a set of links and nodes that represent roads and intersections, respectively. The bridges are assumed to determine the functionality of the link in which they are located. The functionality of the bridges depends on the level of damage reached given a certain earthquake. The point-based earthquake is located at 35.927 N, 89.919W at 10 km below the ground surface.
To start working with this use case, open the Jupyter Notebook on the published project using the button below (same notebook as above).
It may be necessary to click on "Run">"Run All Cells" to allow the visualization of some of the interactive figures. Note: Some cells are used to save figures, which will present an error because the published notebook is in a "Read Only" folder. To run these specific cells or save customized figures, copy the notebook and the input files to your "My Data" folder, as explained below.
Types of visualization tools
This use case focuses on the visualization of static maps, interactive maps, and animated plots. Static plots are a common tool in phases such as model definition and model analysis, where the data needs to be visualized and curated before any calculations, or data publication in formats where the interactivity with the figures does not occur (for example, reports or journal publications). To perform fine explorations of the data collected from an inspection campaign or results from a suite of simulations, it may be necessary to use a more powerful format, such as interactive maps. These are maps in which the user can pan over the map to view data, zoom in or out to examine local or regional groupings, or hover over objects to inspect (initially hidden) information. Finally, animated maps and plots are presented as a way to depict time-dependent geographical information or the distributed influence of feature variations (for example, the increment in ground accelerations as a function of earthquake magnitude). In this case, the user can interact with input features that modify the spatially distributed data or create gifs to show sequential or evolving processes.
Workflow of this use case
The following structure is used:
- Plotting distributed infrastructure using static plots.
- Interactive exploration of spatially distributed information.
- Creation of animated graphs and GIFs.
Setting the python environment
The base image of the Jupyter Notebook DS may (or may not) have pre-installed some of the required Python libraries. The accompanying Jupyter Notebook shows the required libraries in the section: Required installations. Some of the libraries you may need to install include contextily, basemap, plotly, and folium.
Input files
To use the Jupyter Notebook, you need to first create a new folder in your “My Data” and copy the notebook and the input files presented in this published project. The input files have been pre-processed and conveniently shared in the Risk and resilience data examples on DS:
- Hazard:
- CSV files with a mesh grid and bridges’ locations.
- CSV files with the values of the peak ground acceleration (PGA) at the grid and bridge locations for one earthquake event realization (magnitude 7.1).
- Shapefiles: shapefiles of the bridges (points) and roads (polylines).
- Damage results: CSV file with the bridges’ probabilities of reaching or exceeding damage states (obtained using pyincore).
- Network model: TXT files with information on the edges, nodes, and bridge parameters (following the National Bridge Inventory database and additional features).
- Damage_Recovery: CSV and JSON files with the bridges’ probabilities of reaching or exceeding damage states for different magnitude events (obtained using pyincore) and the bridges mean recovery times for a single event.
1. Plotting distributed infrastructure using static plots
Different strategies are used to visualize geographical data easily. In the accompanying Jupyter Notebook libraries such as Geopandas, Matplotlib, Basemap, and Contextily are leveraged to obtain static maps. The examples include visualization of shape files to visualization of networked data (using NetworkX library), both formats in which a user may have the input exposure data. The use case presents how these libraries can be used to create and handle maps and their attributes, such as axis labels, color bars, legends, etc., to highlight features of interest. As shown in Figure 2a, the shapefiles of the roadway network and the bridge locations are shown with the ‘Stamen Terran’ web tiles in the background; the figure also depicts the bridge’s construction year using a ‘Red-Yellow-Blue’ color bar. Figure 2b depicts the case in which resilient modeling outputs such as hazard intensities are used for the background map.
In general, web tiles (available through Python libraries such as Contextily) are used for background maps to enhance the visualization of hazard, exposure, and risk data. Some of these background tiles may require a projection of the coordinates of the data coordinate reference system (CRS) to the CRS of the desired tile, as explained in the Jupyter Notebook.
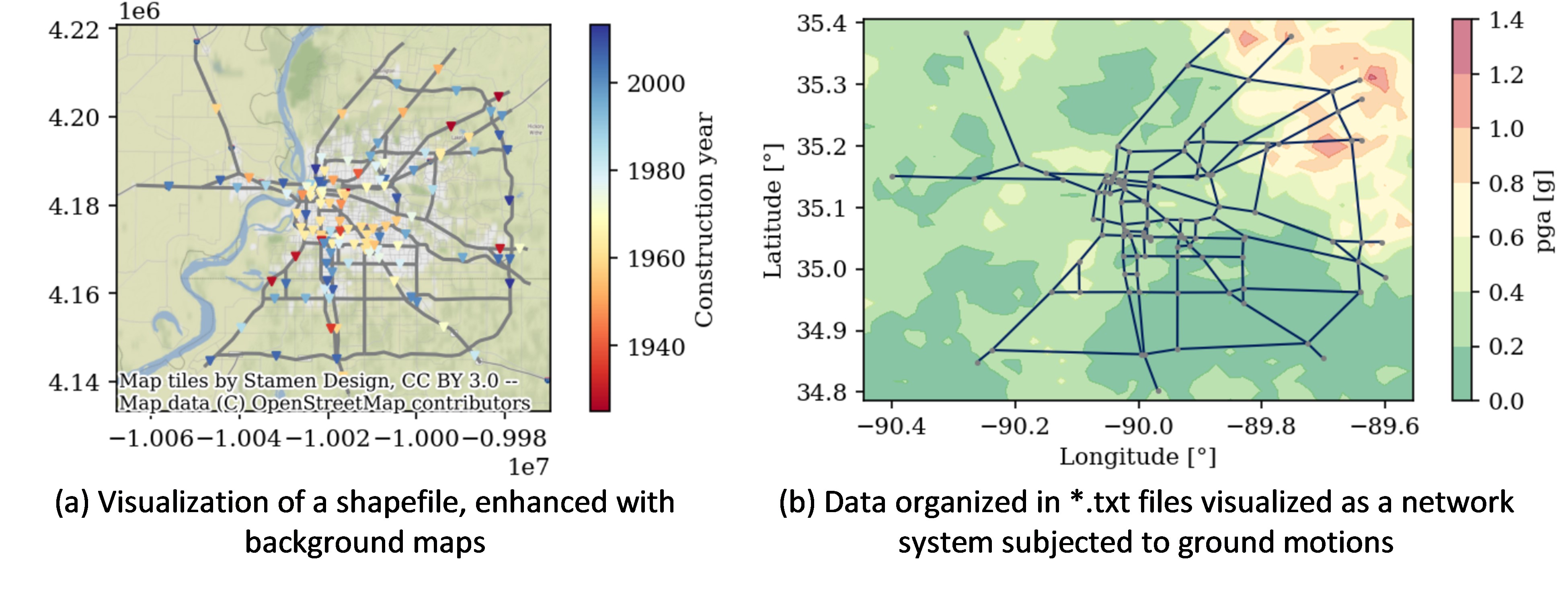
Figure 2. Visualization of static plots
2. Interactive exploration of spatially distributed information
These interactive maps are useful in situations when data visualization is not fully addressed through static maps, so it is necessary to be able to reveal data on elements (points, lines, or polygons) interactively by the user. For example, for inspecting post-event hazard damages, depicting current conditions on situational awareness tools, or displaying information on the infrastructure assets during the restoration processes.
In this use case, damage state exceedance probabilities are obtained for each bridge (i.e., considered as an outcome of applying fragility models to the hazard scenario). An example of this output is presented in the ‘bridge_result.csv’ file for one hazard scenario. In this file format, damage state exceedance probabilities $\mathbb{P}(DS \geq ds_i)$ are named as "$LS_i$", for $i=1,…,4$; similarly, probabilities of being in a damage state $\mathbb{P}(DS = ds_i)$ are named as "$DS_i$", for $i=1,…,4$. For such cases, there may be interest in visualizing the spatial distribution of damage to infrastructure components.
Here, interactive Python libraries are used to visualize and inspect fine information on the different components that comprise the map, such as bridge location, basic information, and damage condition (see Figure 3). These interactive functionalities are integrated using Python libraries such as Plotly and Folium; these allow the user to pan over the different geospatially distributed systems and inspect the region or assets of interest. Also, these enable the user to construct icon objects that display data of interest (e.g. the ‘construction year’ and the ‘exceeding probability of damage state 3’ in Figure 3a) when hovering over the bridge locations. If additional data is also important to display (e.g. hazard intensity, link, or bridge IDs, among others), ‘pop-up’ functionalities can be used to present this information when the user clicks on a particular object (shown in Figure 3b).
As shown in this use case, interactive maps can be enhanced by handling the icons, points, and link characteristics such as type, icon figure, color, etc. Figure 3 presents the bridge condition using a common color coding related to post-hazard tagging. Red tag is used here when $\mathbb{P}(DS≥ds_3 )≥0.15$, yellow tag is used if $0.05≤\mathbb{P}(DS≥ds_3 )<0.15$, and green tag is used if $\mathbb{P}(DS≥ds_3)<0.05$; note that these limits have been arbitrarily selected for display purposes. Moreover, objects such as legends and color bars can be easily included in such interactive maps to add additional layers of information. Given the possibility of presenting the data "online", these are very useful tools for communication with stakeholders, inspection teams, or simply for data analysis during damage simulation or recovery processes.
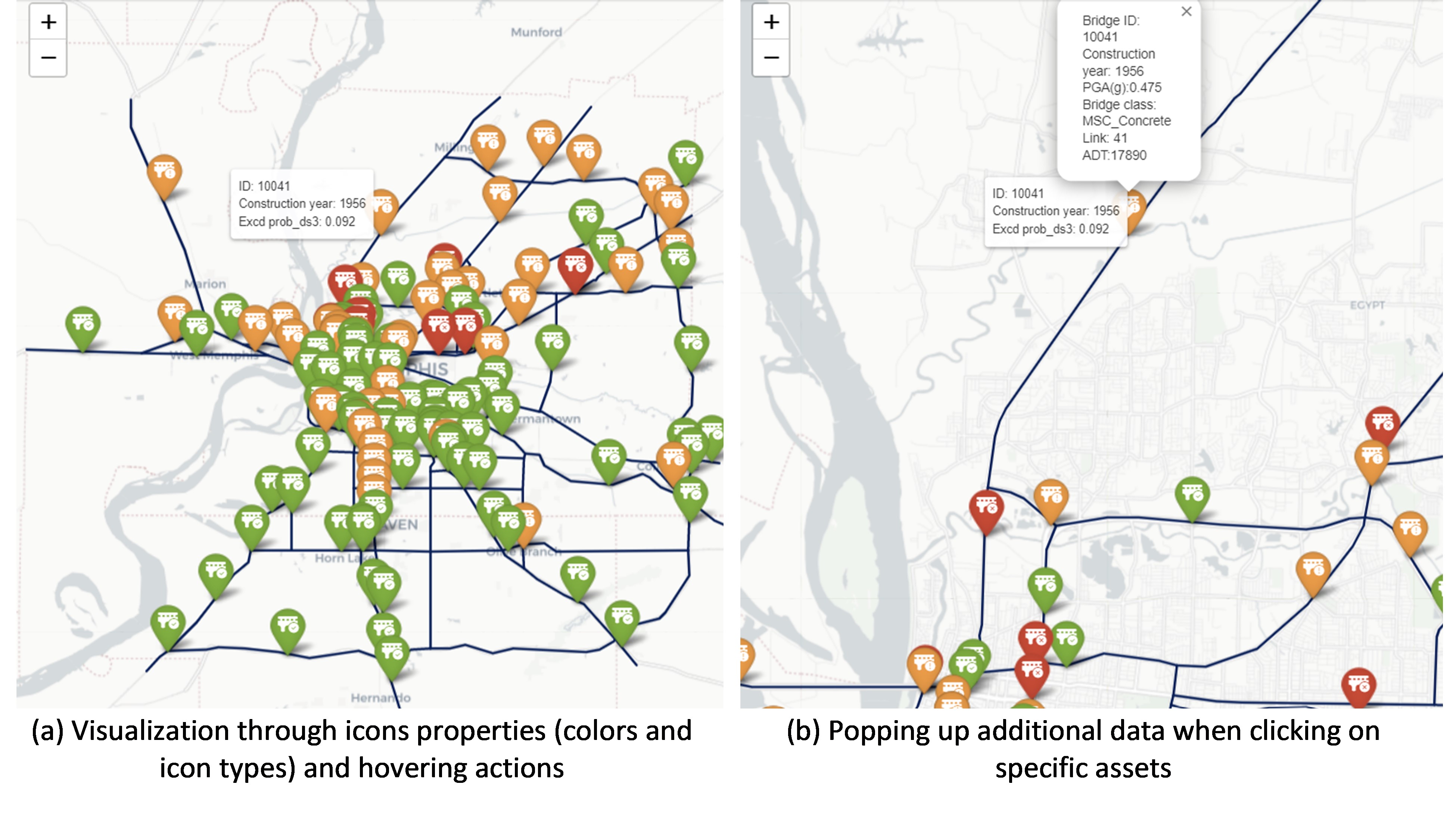
Figure 3. Visualization of interactive plots
3. Creation of animated graphs and GIFs
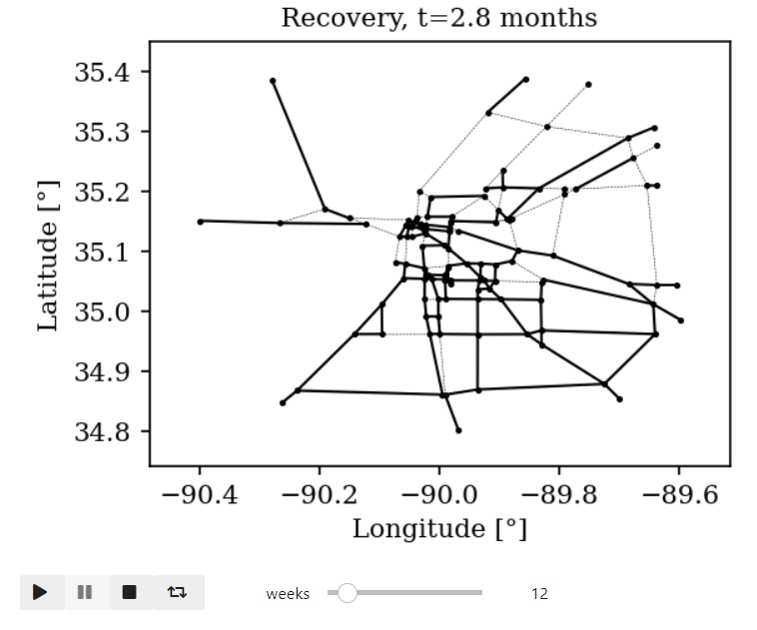
Time-varying characteristics or feature-dependent results can be effectively visualized and communicated using animations that enable the user to modify certain independent variables such as the magnitude of an earthquake, the number of crews used to repair a networked system, or the level of acceptable damage before demanding evacuation. Common sources of time-dependent data in resilient modeling may include the status of components as repair and recovery processes evolve. In this use case, the highway functionality is visualized using solid lines if the link is ‘fully functional’, or through dashed lines if any of the bridges on the route are ‘under repair’ (see Figure 4).
Different widgets (such as checkboxes, sliders, or buttons) can be passed to the Jupyter Notebook using Ipywidgets packages to create animations that facilitate user-graphics interaction. These animations allow the user to select specific characteristics (or a combination of these) to visualize and interact with the data. For example, Figure 4a shows how the user can skip forward or backward in time (weeks) or drag the slider to the point in time in which it is desired to know the network state. Hence, the buttons enable the creation of dynamic figures that sequentially vary the features of interest, depicting their influence on the distributed data. These animated ‘videos’ can also be presented as GIFs (i.e., graphics interchange format), as shown in Figure 4b, which are easy to visualize and share across different platforms or programs, for example, into phone applications, slides, reports or webpages.
| (a) Example of a graph animated with interactive widgets | (b) Example of a ‘gif’ animation |
|---|---|
 |
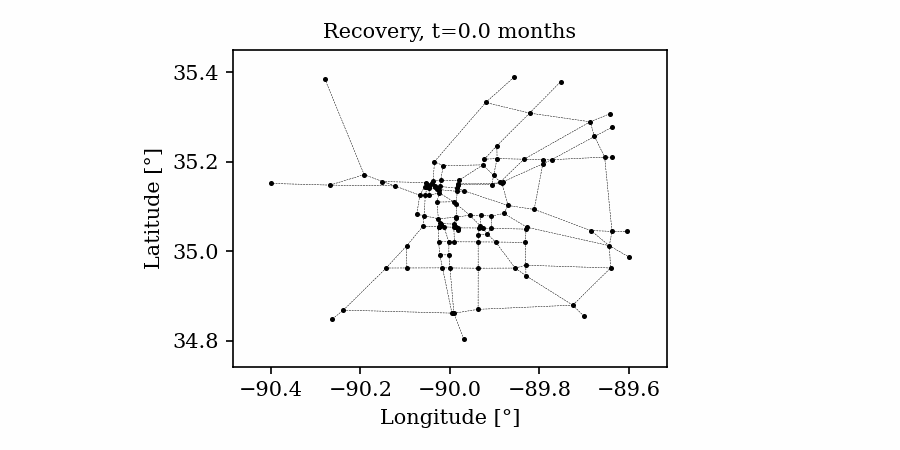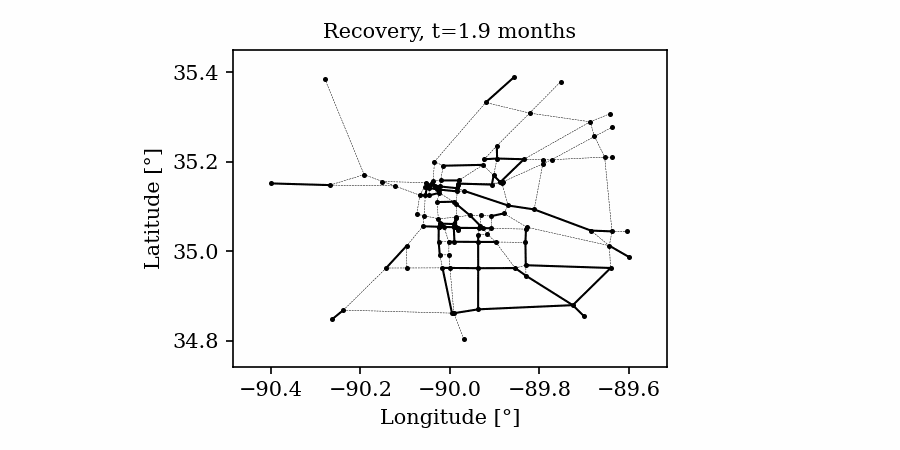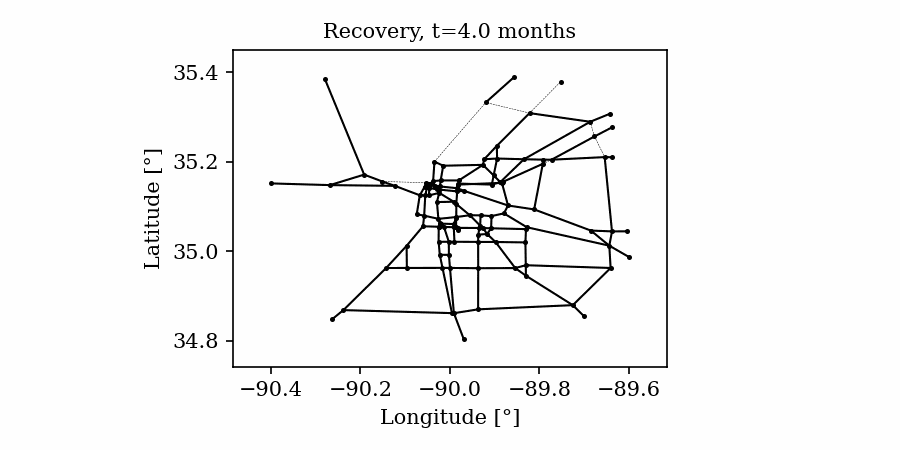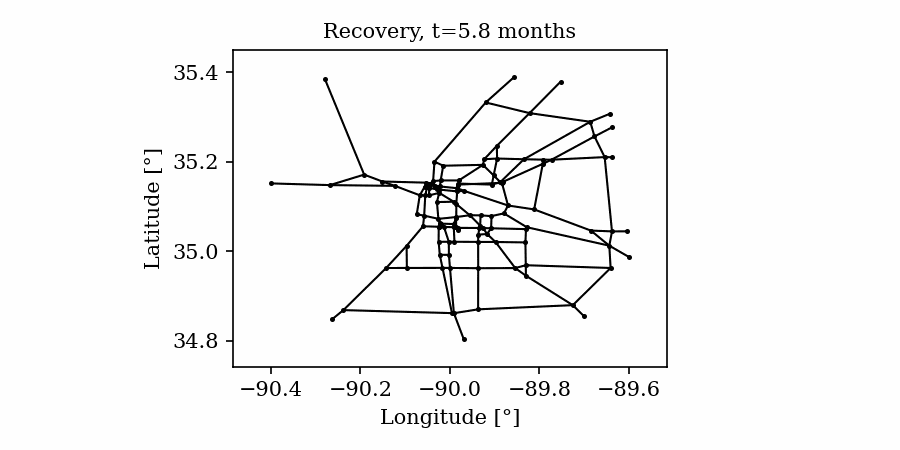 |
Figure 4. Animated plots for visualization of dynamic outputs.
Geohazards
- Data Analysis using Next Generation Liquefaction (NGL) Database (NGL, Jupyter, SQL, Interactive Data Analytics)
- Landslide Runout Simulations (MPM, Jupyter, Paraview, HPC)
NGL Database
NGL Database
Next Generation Liquefaction (NGL) Database Jupyter Notebooks
Brandenberg, S.J. - UCLA
Ulmer, K.J. - Southwest Research Institute
Zimmaro, P. - University of Calabria
The example makes use of the following DesignSafe resources:
Jupyter notebooks on DS Juypterhub
NGL Database
Background
Citations and Licensing
-
Please cite Zimmaro, P., et al. (2019) to acknowledge the use of the NGL Database. Data in the NGL database has been gathered from these published sources. If you use specific data in the database, please cite the original source.
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License.
Description
The Next Generation Liquefaction (NGL) Project is advancing the state of the art in liquefaction research and working toward providing end users with a consensus approach to assess liquefaction potential within a probabilistic and risk-informed framework. Specifically, NGL’s goal is to first collect and organize liquefaction information in a common and comprehensive database to provide all researchers with a substantially larger, more consistent, and more reliable source of liquefaction data than existed previously. Based on this database, we will create probabilistic models that provide hazard- and risk-consistent bases for assessing liquefaction susceptibility, the potential for liquefaction to be triggered in susceptible soils, and the likely consequences. NGL is committed to an open and objective evaluation and integration of data, models and methods, as recommended in a 2016 National Academies report.
The evaluation and integration of the data into new models and methods will be clear and transparent. Following these principles will ensure that the resulting liquefaction susceptibility, triggering, and consequence models are reliable, robust and vetted by the scientific community, providing a solid foundation for designing, constructing and overseeing critical infrastructure projects.
The NGL database is populated through a web interface at www.nextgenerationliquefaction.org/. The web interface provides limited capabilities for users to interact with data. Users are able to view and download data, but they cannot use the GUI to develop an end-to-end workflow to make scientific inferences and draw conclusions from the data. To facilitate end-to-end workflows, the NGL database is replicated daily to DesignSafe, where users can interact with it using Jupyter notebooks.
Understanding the Database Schema
The NGL database is organized into tables that are related to each other via keys. To query the database, you will need to understand the organizational structure of the database, called the schema. The database schema is documented at the following URL:
https://nextgenerationliquefaction.org/schema/index.html
Querying Data via Jupyter Notebooks
Jupyter notebooks provide the capability to query NGL data, and subsequently process, visualize, and learn from the data in an end-to-end workflow. Jupyter notebooks run in the cloud on DesignSafe, and provide a number of benefits compared with a more traditional local mode of operation:
- The NGL database contains many GB of data, and interating with it in the cloud does not require downloading these data files to a local file system.
- Users can collaborate in the cloud by creating DesignSafe projects where they can share processing scripts.
- The NGL database is constantly changing as new data is added. Working in the cloud means that the data will always be up-to-date.
- Querying the MySQL database is faster than opening individual text files to extract data.
This documentation first demonstrates how to install the database connection script, followed by several example scripts intended to serve as starting points for users who wish to develop their own tools.
Installing Database Connection Script
Connecting to a relational database requires credentials, like username, password, database name, and hostname. Rather than requiring users to know these credentials and create their own database connections, we have created a Python package that allows users to query the database. This code installs the package containing the database connection script for NGL:
!pip install git+https://github.com/sjbrandenberg/designsafe_dbExample Queries
This notebook contains example queries to illustrate how to extract data from the NGL database into Pandas dataframe objects using Python scripts in Jupyter notebooks. The notebook contains cells that perform the following operations:
- Query contents of the SITE table
- Query event information and associated field observations at the Wildlife liquefaction array
- Query cone penetration test data at Wildlife liquefaction array
- Query a list of tables in the NGL database
- Query information about BORH table
- Query counts of cone penetration test data, boreholes, surface wave measurements, invasive shear wave velocity measurements, liquefaction observations, and non-liquefaction observations
Cone Penetration Test Viewer
The cone penetration test viewer demonstrates the following:
- Connecting to NGL database in DesignSafe
- Querying data from SITE, TEST, SCPG, and SCPT tables into Pandas dataframes
- Creating dropdown widgets using the ipywidgets package to allow users to select site and test data
- Creating HTML widget for displaying metadata after a user select a test
- Using the ipywidgets "observe" feature to call functions when users select a widget value
- Plotting data from the selected cone penetration test using matplotlib
Cone penetration test data plotted in the notebook include tip resistance, sleeve friction, and pore pressure. In some cases, sleeve friction and pore pressure are not measured, in which case the plots are empty.
VS (Invasive) Test Viewer
The Vs (Invasive) Test Viewer demonstrates the following:
- Connecting to NGL database in DesignSafe
- Querying data from SITE, TEST, GINV, and GIND tables into Pandas dataframes
- Creating dropdown widgets using the ipywidgets package to allow users to select site and test data
- Creating HTML widget for displaying metadata after a user selects a test
- Using the ipywidgets "observe" feature to call functions when users select a widget value
- Plotting data from the selected invasive geophysical test using matplotlib
October 2021 DesignSafe Webinar
The DesignSafe_Webinar_Oct2021 notebook was created during a webinar/workshop hosted by DesignSafe and the Pacific Earthquake Engineering Research (PEER) center.
The notebook demonstrates the following:
- Connecting to NGL database in DesignSafe
- Querying data from SITE, TEST, SCPG, and SCPT tables into Pandas dataframes
- Plotting data from the selected test using matplotlib
Cone penetration test data plotted in the notebook include tip resistance, sleeve friction, and pore pressure. In some cases, sleeve friction and pore pressure are not measured, in which case the plots are empty.
DesignSafe_Webinar_Oct2021.ipynb
DesignSafe Webinar YouTube video
DesignSafe Workshop YouTube video
Direct Simple Shear Laboratory Test Viewer
The Direct Simple Shear Laboratory Test Viewer is a graphical interface that plots direct simple shear tests in the NGL database. It demonstrates the following:
- Connecting to NGL database in DesignSafe
- Querying data from LAB, LAB_PROGRAM, SAMP, SPEC, DSSG, and DSSS tables into Pandas dataframes
- Creating dropdown widgets using the ipywidgets package to allow users to select lab, sample, specimen, and test data
- Creating javascript for downloading the selected direct simple shear test to a local computer
- Plotting data from the selected direct simple shear test using matplotlib
Direct simple shear data plotted in the notebook include shear stress, shear strain, vertical stress, and vertical strain time series in the first plot. The second plot displays shear strain and void ratio versus vertical stress and void ratio, shear stress, and vertical stress ratio versus shear strain.
MPM Landslide
MPM Landslide
Material Point Method for Landslide Modeling
Krishna Kumar - University of Texas at Austin
The example makes use of the following DesignSafe resources:
Jupyter notebooks on DS Juypterhub
CB-Geo MPM
ParaView
Background
Citation and Licensing
- Please cite Kumar et al. (2019) to acknowledge the use of CB-Geo MPM.
- Please cite Abram et al. (2022) to acknowledge the use of any resources from the Oso in situ use case.
- Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
- This software is distributed under the MIT License.
Description
Material Point Method (MPM) is a particle based method that represents the material as a collection of material points, and their deformations are determined by Newton’s laws of motion. The MPM is a hybrid Eulerian-Lagrangian approach, which uses moving material points and computational nodes on a background mesh. This approach is very effective particularly in the context of large deformations.
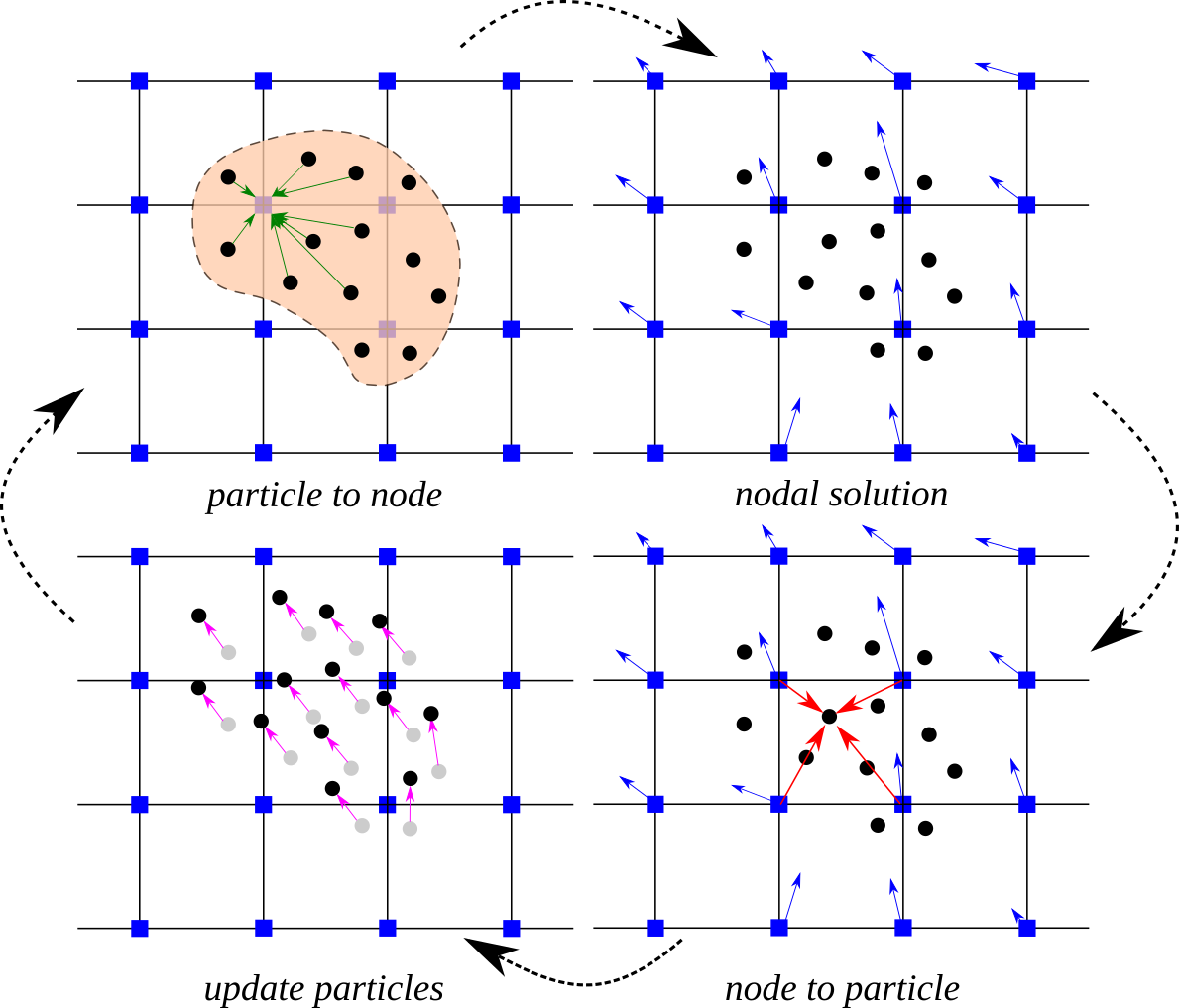
Illustration of the MPM algorithm (1) A representation of material points overlaid on a computational grid. Arrows represent material point state vectors (mass, volume, velocity, etc.) being projected to the nodes of the computational grid. (2) The equations of motion are solved onto the nodes, resulting in updated nodal velocities and positions. (3) The updated nodal kinematics are interpolated back to the material points. (4) The state of the material points is updated, and the computational grid is reset.
This use case demonstrates how to run MPM simulations on DesignSafe using Jupyter Notebook. For more information on CB-Geo MPM visit the GitHub repo and user documentation.
Input generation
Input files for the MPM code can be generated using pycbg. The documentation of the input generator is here. For more information on the input files, please refer to CB-Geo MPM documentation. The generator is available at PyPI and an be easily installed with pip install pycbg. pycbg enables a Python generation of expected .json input files, offering all Python capabilities to CB-Geo MPM users for this preprocessing stage.
Typing a few Python lines is usually enough for a user to define all necessary ingredients for a MPM simulation:
-
generate the mesh (using gmsh)
-
generate the particles
-
define the entity sets
-
create boundary conditions
-
set the analysis' parameters
-
setup batch of simulations (the documentation doesn't mention it yet but the function
pycbg.preprocessing.setup_batchhas a complete docstring)
An example
Simulation of a settling column made with two different materials is described in preprocess.ipynb as follows:
import pycbg.preprocessing as utl
### The usual start of a PyCBG script:
sim = utl.Simulation(title="Two_materials_column")
### Creating the mesh:
sim.create_mesh(dimensions=(1.,1.,10.), ncells=(1,1,10))
### Creating Material Points, could have been done by filling an array manually:
sim.create_particles(npart_perdim_percell=1)
### Creating entity sets (the 2 materials), using lambda functions:
sim.init_entity_sets()
lower_particles = sim.entity_sets.create_set(lambda x,y,z: z<5, typ="particle")
upper_particles = sim.entity_sets.create_set(lambda x,y,z: z>=5, typ="particle")
### The materials properties:
sim.materials.create_MohrCoulomb3D(pset_id=lower_particles)
sim.materials.create_Newtonian3D(pset_id=upper_particles)
### Boundary conditions on nodes entity sets (blocked displacements):
walls = []
walls.append([sim.entity_sets.create_set(lambda x,y,z: x==lim, typ="node") for lim in [0, sim.mesh.l0]])
walls.append([sim.entity_sets.create_set(lambda x,y,z: y==lim, typ="node") for lim in [0, sim.mesh.l1]])
walls.append([sim.entity_sets.create_set(lambda x,y,z: z==lim, typ="node") for lim in [0, sim.mesh.l2]])
for direction, sets in enumerate(walls): _ = [sim.add_velocity_condition(direction, 0., es) for es in sets]
### Other simulation parameters (gravity, number of iterations, time step, ..):
sim.set_gravity([0,0,-9.81])
nsteps = 1.5e5
sim.set_analysis_parameters(dt=1e-3, nsteps=nsteps, output_step_interval=nsteps/100)
### Save user defined parameters to be reused at the postprocessing stage:
sim.add_custom_parameters({"lower_particles": lower_particles,
"upper_particles": upper_particles,
"walls": walls})
### Final generation of input files:
sim.write_input_file()This creates in the working directory a folder Two_materials_column where all the necessary input files are located.
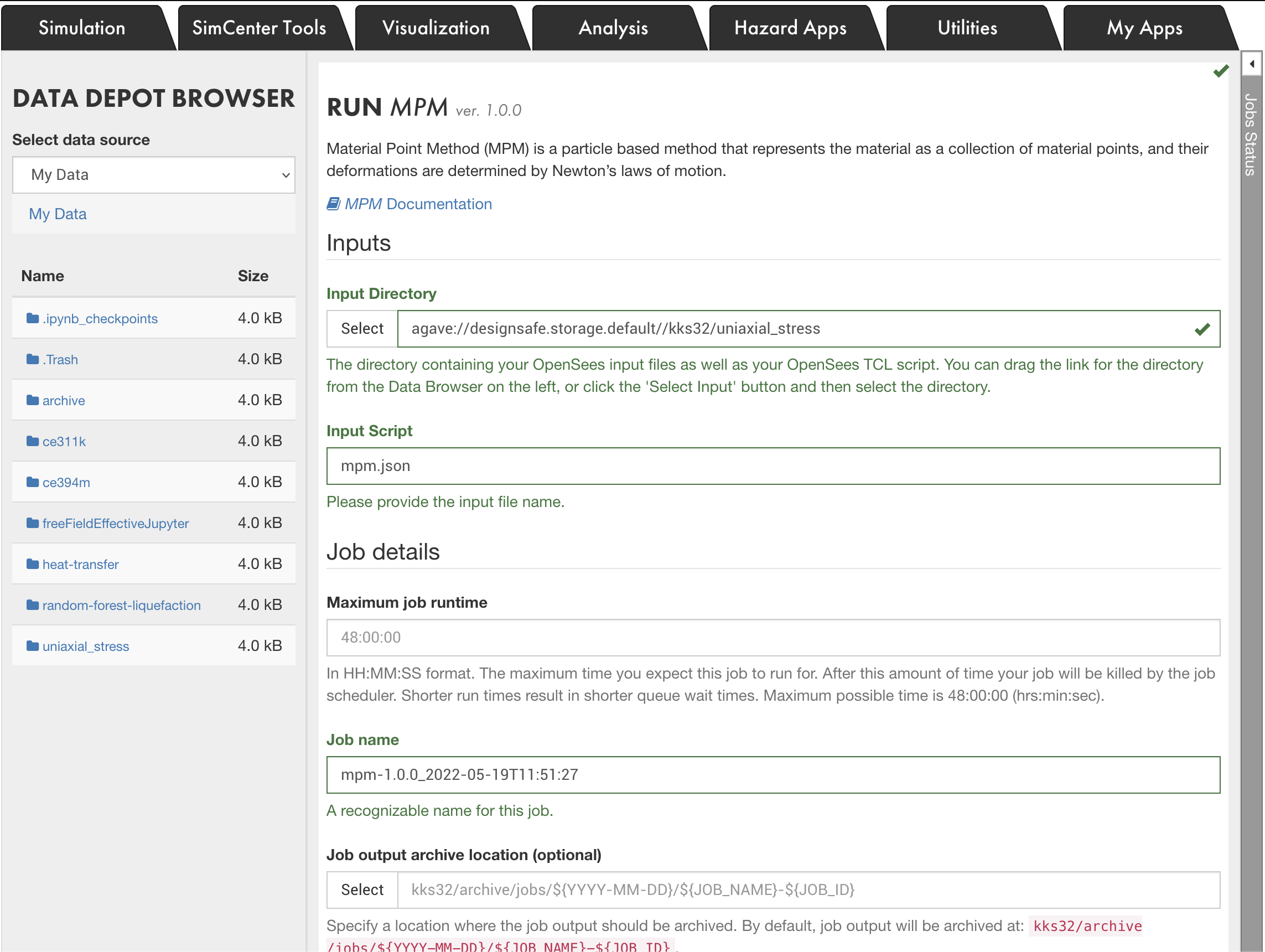
Running the MPM Code
The CB-Geo MPM code is available on DesignSafe under WorkSpace > Tools & Applications > Simulations. Launch a new MPM Job. The input folder should have all the scripts, mesh and particle files. CB-Geo MPM can run on multi-nodes and has been tested to run on upto 15,000 cores.
Post Processing
VTK and ParaView
The MPM code can be set to write VTK data of particles at a specified output frequency. The input JSON configuration takes as optional vtk argument. The following attributes are valid options for VTK: "stresses, strains, and velocities. When the attribute vtk is not specified or an incorrect argument is defined, the code will write all available options.
"post_processing": {
"output_steps": 5,
"path": "results/",
"vtk" : ["stresses","velocities"],
"vtk_statevars": [
{
"phase_id": 0,
"statevars" : ["pdstrain"]
}
]
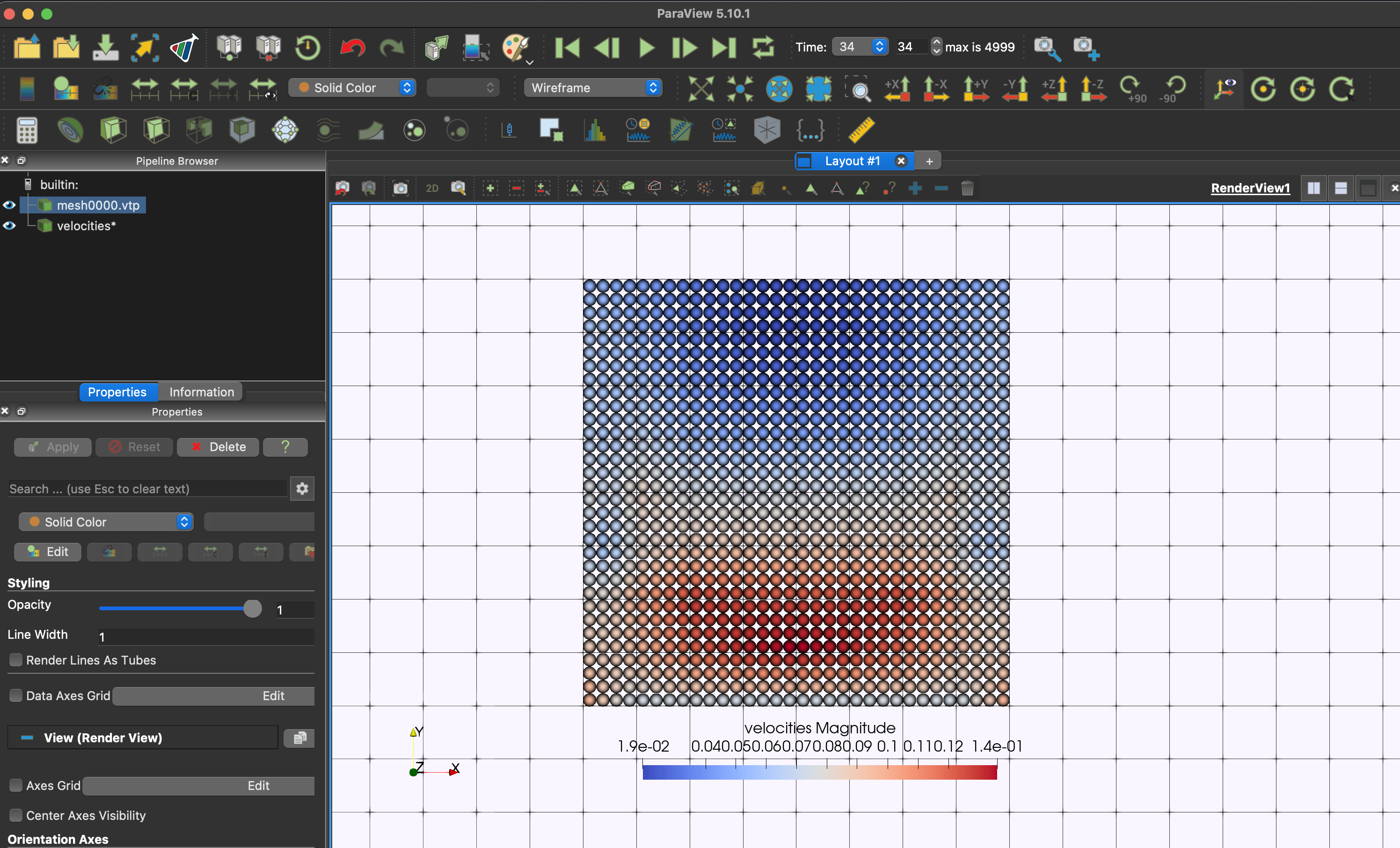
}When opening particle data (*.vtp) in ParaView, please use the representation
Point Gaussianto visualise the particle data attribute.
The CB-Geo MPM code generates parallel *.pvtp files when the code is executed across MPI ranks. Each MPI rank will produce an attribute subdomain files, for example stresses-0_2-100.vtp and stresses-1_2-100.vtp file for stresses generated in rank 0 of 2 rank MPI processes and also a parallel pvtp file stresses-100.pvtp. The parallel *.pvtp file combines all the VTK outputs from different MPI ranks.
Use the
*.pvtpfiles for visualizing results from a distributed simulation. No need to load individual subdomain*.vtpwhen visualizing results from the MPI tasks.
The parameter vtk_statevars is an optional VTK output, which will print the value of the state variable for the particle. If the particle does not have the specified state variable, it will be set to NaN.
You can view the results in DesignSafe ParaView
HDF5
The CB-Geo mpm code writes HDF5 data of particles at each output time step. The HDF5 data can be read using Python / Pandas. If pandas package is not installed, run pip3 install pandas. The postprocess.ipynb shows how to perform data analysis using HDF5 data.
To read a particles HDF5 data, for example particles00.h5 at step 0:
### Read HDF5 data
### !pip3 install pandas
import pandas as pd
df = pd.read_hdf('particles00.h5', 'table')
### Print column headers
print(list(df))The particles HDF5 data has the following variables stored in the dataframe:
['id', 'coord_x', 'coord_y', 'coord_z', 'velocity_x', 'velocity_y', 'velocity_z',
'stress_xx', 'stress_yy', 'stress_zz', 'tau_xy', 'tau_yz', 'tau_xz',
'strain_xx', 'strain_yy', 'strain_zz', 'gamma_xy', 'gamma_yz', 'gamma_xz', 'epsilon_v', 'status']Each item in the header can be used to access data in the h5 file. To print velocities (x, y, and z) of the particles:
### Print all velocities
print(df[['velocity_x', 'velocity_y','velocity_z']]) velocity_x velocity_y velocity_z
0 0.0 0.0 0.016667
1 0.0 0.0 0.016667
2 0.0 0.0 0.016667
3 0.0 0.0 0.016667
4 0.0 0.0 0.033333
5 0.0 0.0 0.033333
6 0.0 0.0 0.033333
7 0.0 0.0 0.033333Oso landslide with in situ visualization
In situ visualization is a broad approach to processing simulation data in real-time - that is, wall-clock time, as the simulation is running. Generally, the approach is to provide data extracts, which are condensed representations of the data chosen for the explicit purpose of visualization and computed without writing data to external storage. Since these extracts (often images) are vastly smaller than the raw simulation itself, it becomes possible to save them at a far higher temporal frequency than is practical for the raw data, resulting in substantial gains in both efficiency and accuracy. In situ visualization allows simulations to export complete datasets only at the temporal frequency necessary for economic check- point/restart.
We leverage in situ viz with MPM using TACC Galaxy.
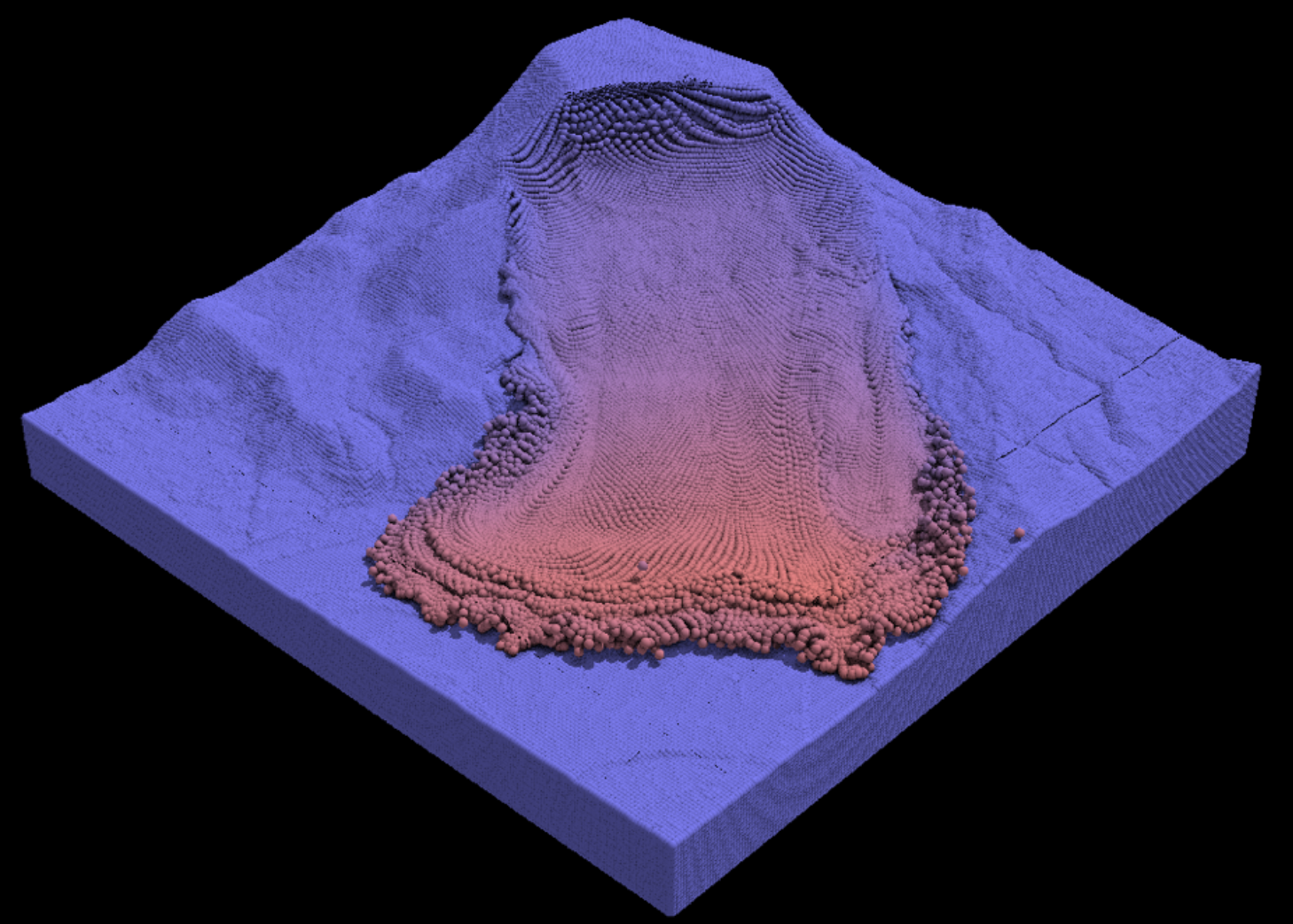

In situ rendering of the Oso landslide with CB-Geo MPM of 5 million material points running 16 MPI tasks for compute + 8 MPI tasks for visualization.
Wind & Storm Surge
- Large-Scale Ensemble Simulations of Storm Surge (ADCIRC, pylauncher, Jupyter, HPC)
- Creating an ADCIRC dataset (ADCIRC, pylauncher, Jupyter, Dataset)
- Visualization of Storm Surge Impacts (ADCIRC, Jupyter, QGIS)
- Simulation of Wind Flow around Buildings (OpenFOAM, Jupyter, Paraview, HPC)
- Analysis of Field Sensor Data from Wind Events (Jupyter, Interactive Data Analysis)
Large-Scale Storm Surge
Large-Scale Storm Surge
ADCIRC Use Case - Using Tapis and Pylauncher for Ensemble Modeling in DesignSafe
Clint Dawson, University of Texas at Austin
Carlos del-Castillo-Negrete, University of Texas at Austin
Benjamin Pachev, University of Texas at Austin
The following use case presents an example of how to leverage the Tapis API to run an ensemble of HPC simulations. The specific workflow to be presented consists of running ADCIRC, a storm-surge modeling application available on DesignSafe, using the parametric job launcher pylauncher. All code and examples presented are meant to be be executed from a Jupyter Notebook on the DesignSafe platform and using a DesignSafe account to make Tapis API calls.
Resources
Jupyter Notebooks
Accompanying jupyter notebooks for this use case can be found in the ADCIRC folder in Community Data. You may access these notebooksdirectly:
| Scope | Notebook |
|---|---|
| Create an ADCIRC DataSet | Creating an ADCIRC DataSet.ipynb |
| Create an Ensemble Simulations | ADCIRC Ensemble Simulations.ipynb |
DesignSafe Resources
The following DesignSafe resources were used in developing this use case.
Background
Citation and Licensing
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License.
ADCIRC
For more information on running ADCIRC and documentation, see the following links:
ADCIRC is available as a standalone app accesible via the DesignSafe front-end.
Tapis
Tapis is the main API to control and access HPC resources with. For more resources and tutorials on how to use Tapis, see the following:
To initialize tapis in our jupyter notebook we use AgavePy. Relies on tapis auth init --interactive being run from a terminal first upon initializing your Jupyter server.
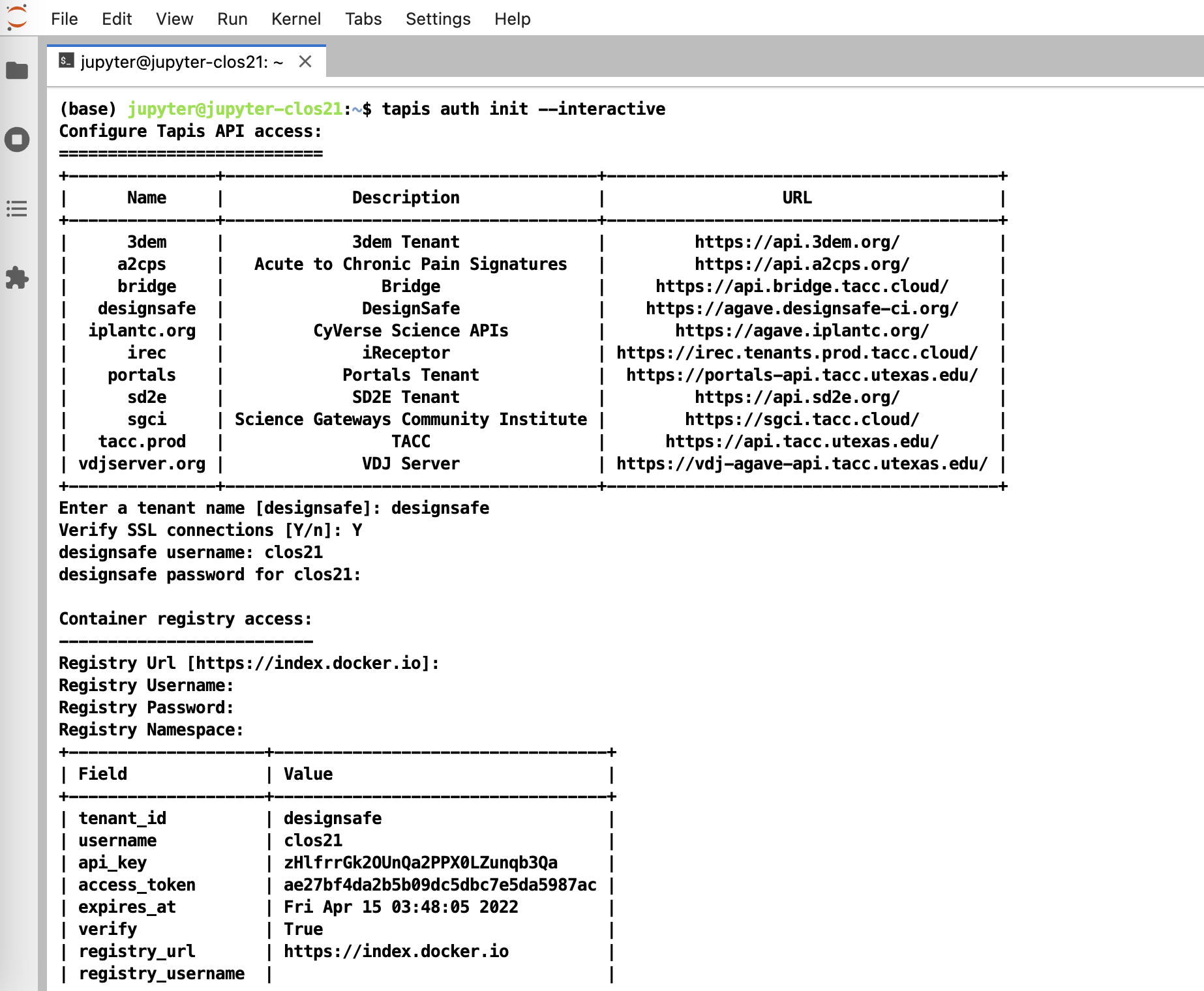
Initialize Tapis from within a shell in a jupyter session. A shell can be launched by going to File -> New -> Terminal.
Once this is complete, you can now run from a code cell in your jupyter session the following to load your AgavePy credentials:
from agavepy.agave import Agave
ag = Agave.restore()Pylauncher
Pylauncher is a parametric job launcher used for launching a collection of HPC jobs within one HPC job. By specifying a list of jobs to execute in either a CSV or json file, pylauncher manages resources on a given HPC job to execute all the jobs using the given nodes. Inputs for pylauncher look something like (for csv files, per line):
num_processes,<pre process command>;<main parallel command>;<post process command>The pre-process and post-process commands are executed in serial, while the main command is executed in parallel using the appropriate number of processes. Note pre and post process commands should do light file management and movement and no computationally intensive tasks.
Tapis Pylauncher App
Overview of this section:
- Getting the Appication
- App Overview
- Staging Files
- Example Ensemble ADCIRC RUN
Accessing the Application
The code for the tapis application is publicly accessible at https://github.com/UT-CHG/tapis-pylauncher. A public Tapis application exists using version 0.0.0 of the application deployed under the ID pylauncher-0.0.0u1.
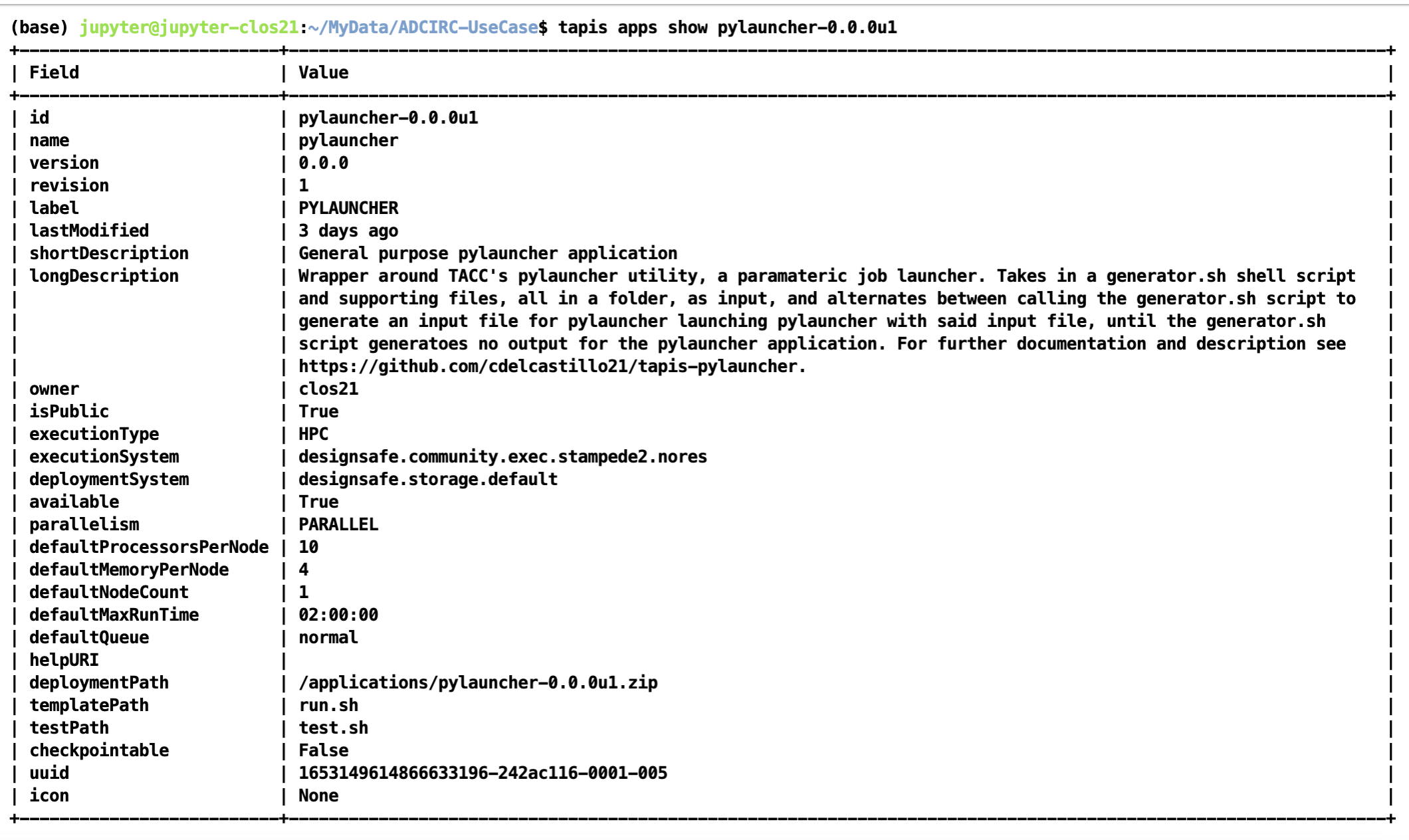
The publicly available pylauncher application should be available to all users via the CLI/API, but will not be visible via DesignSafe's workspaces front-end.
Basic Application Overview
The tapis-pylauncher application loops through iterations of calling pylauncher utility, using as input a file generated by a user defined generator shell script generator.sh. A simplified excerpt of this main execution loop is as follows:
### Main Execution Loop:
### - Call generator script.
### - Calls pylauncher on generated input file. Expected name = jobs_list.csv
### - Repeats until generator script returns no input file for pylauncher.
ITER=1
while :
do
# Call generator if it exists script
if [ -e generator.sh ]
then
./generator.sh ${ITER} $SLURM_NPROCS $generator_args
fi
# If input file for pylauncher has been generated, then start pylauncher
if [ -e ${pylauncher_input} ]
then
python3 launch.py ${pylauncher_input} >> pylauncher.log
fi
ITER=$(( $ITER + 1 ))
doneNote how a generator script is not required, with a static pylauncher file, of input name determined as a job parameter pylauncher_input, being sufficient to run a single batch of jobs.
All input scripts and files for each parametric job should be zipped into a file and passed as an input to the pylauncher application. Note that these files shouldn't be too large and shouldn't contain data as tapis will be copying them around to stage and archive jobs. Data should ideally be pre-staged and not part of the zipped job inputs.
Staging Files
For large scale ensemble simulations, it is best to stage individual ADCIRC run files in a project directory that execution systems can access before-hand so that Tapis itself isn't doing the moving and staging of data.
The corresponding TACC base path to your project with a particular id can be found at /corral-repl/projects/NHERI/projects/[id]/. To find the ID for your project, you can just look at the URL of your project directory in designsafe:
TX FEMA storms project directory. Note how the URL on top contains the Project ID corresponding to the path on corral that login nodes on TACC systems should have access to.
From a login node then (assuming this is done on stampede2), the data can be staged onto a public directory on /work as follows. First we create a public directory in our workspace where the data will be staged:
(base) login2.stampede2(1020)$ cd $WORK
(base) login2.stampede2(1022)$ cd ..
(base) login2.stampede2(1023)$ ls
frontera lonestar longhorn ls6 maverick2 pub singularity_cache stampede2
(base) login2.stampede2(1024)$ pwd
/work2/06307/clos21
(base) login2.stampede2(1026)$ chmod o+x
(base) login2.stampede2(1027)$ mkdir -p pub
(base) login2.stampede2(1028)$ chmod o+x pub
(base) login2.stampede2(1029)$ cd pub
(base) login2.stampede2(1030)$ mkdir -p adcirc/inputs/ShinnecockInlet/mesh/testNext we copy the data from our project directory to the public work directory
(base) login2.stampede2(1039)$ cp /corral-repl/projects/NHERI/projects/586025607221276180-242ac118-0001-012/ShinnecockInletMesh/* adcirc/inputs/ShinnecockInlet/mesh/test/Finally we change the ownership of the files and all sub-directories where the data is staged to be publicly accessible by the TACC execution systems. Which we can check via the file permissions of the final directory we created with the staged data:
(base) login2.stampede2(1040)$ chmod -R a-x+rX adcirc
(base) login2.stampede2(1042)$ cd adcirc/inputs/ShinnecockInlet/mesh/test
(base) login2.stampede2(1043)$ pwd
/work2/06307/clos21/pub/adcirc/inputs/ShinnecockInlet/mesh/test
(base) login2.stampede2(1045)$ ls -lat
total 360
-rw-r--r-- 1 clos21 G-800588 341496 May 13 17:27 fort.14
-rw-r--r-- 1 clos21 G-800588 15338 May 13 17:27 fort.15
drwxr-xr-x 2 clos21 G-800588 4096 May 13 17:26 .
drwxr-xr-x 4 clos21 G-800588 4096 May 13 17:24 ..The directory /work2/06307/clos21/pub/adcirc/inputs/ShinnecockInlet/mesh/test now becomes the directory we can use in our pylauncher configurations and scripts to access the data to be used for the ensemble simulations.
Example Ensemble Run: Shinnecock Inlet Test Grid Performance
For an example of how to use the tapis-pylauncher application, we refer to the accompanying notebook in the ADCIRC Use Case folder in the Community Data directory.
The notebook goes over how to run ADCIRC on the Shinnecock Inlet Test Grid.
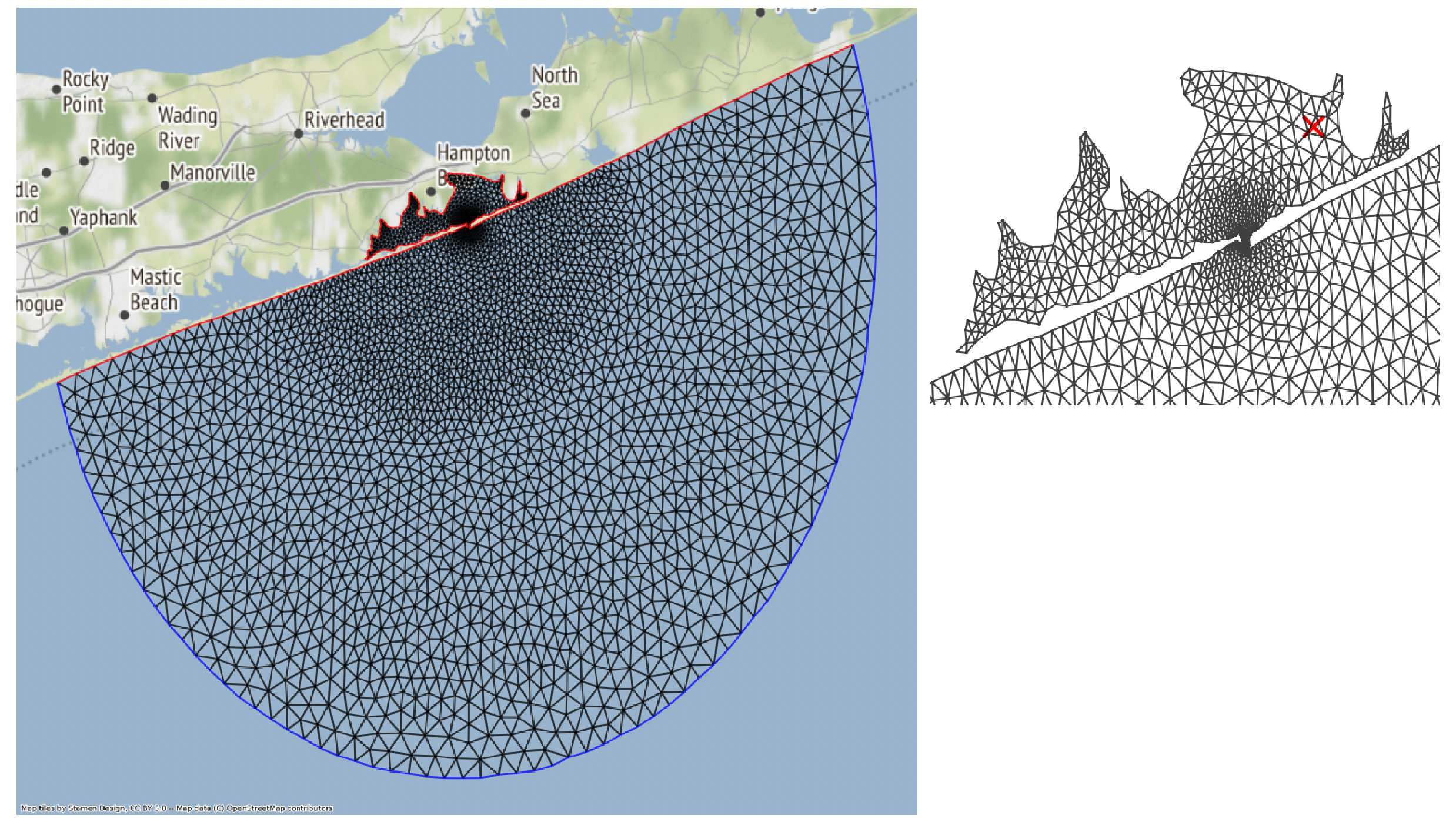
Shinnecock Inlet Test Grid. ADCIRC solves the Shallow Water Equations over a Triangular Mesh, depicted above.
An ensemble of adcirc simulations using different amounts of parallel processes on the same grid is configured, and output from active and archived job runs is analyzed to produced bar plots of run-time versus number of processors used for the Shinneocock Inlet Grid.
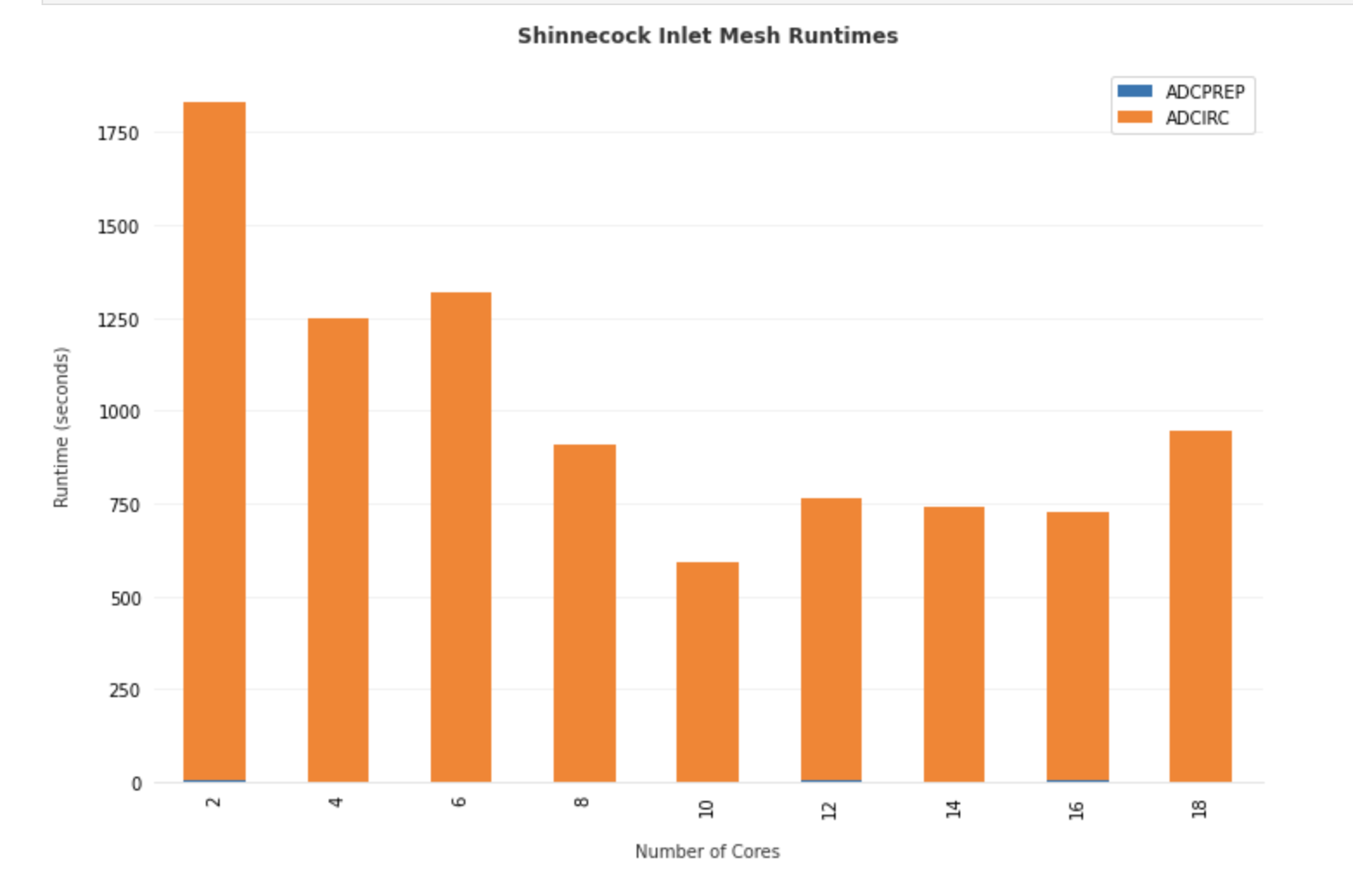
Total Runtime for ADCIRC on the Shinnecock Inlet grid pictured above using different number of processors on stampede2.
Creating an ADCIRC dataset
ADCIRC Datasets
ADCIRC Use Case - Creating an ADCIRC DataSet on DesignSafe
Clint Dawson, University of Texas at Austin
Carlos del-Castillo-Negrete, University of Texas at Austin
Benjamin Pachev, University of Texas at Austin
Overview
The following use case demonstrates how to compile an ADCIRC data-set of hind-casts on DesignSafe. This workflow involves the following steps:
- Finding storm-surge events.
- Compiling meteorological forcing for storm surge events.
- Running ADCIRC hind-casts using meteorological forcing.
- Organize and publish data on DesignSafe, obtaining a DOI for your research and for others to cite your data when re-used.
The workflow presented here is a common one performed for compiling ADCIRC data-sets for a variety of purposes, from Uncertainty Quantification to training Surrogate Models. Whatever your application is of ADCIRC data, publishing your dataset on DesignSafe allows you to re-use your own data, and for others to use and cite your data as well.
To see a couple of Example data-sets, and associated published research using the datasets, see the following examples:
- Texas FEMA Storms - Synthetic storms for assessing storm surge risk. Used recently in Pachev et. al 2023 to train a surrogate model for ADCIRC for the coast of Texas.
- Alaska Storm Surge Events - Major storm surge events for the coast of Alaska. Also used in Pachev et. al 2023 for creating a surrogate model for the coast of Alaska.
An accompanying jupyter notebook for this use case can be found in the ADCIRC folder in Community Data under the name Creating an ADCIRC DataSet.ipynb.
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
| Scope | Notebook |
|---|---|
| Create an ADCIRC DataSet | Creating an ADCIRC DataSet.ipynb |
DesignSafe Resources
The following DesignSafe resources were leveraged in developing this use case.
Background
Citation and Licensing
- Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
- This software is distributed under the GNU General Public License.
ADCIRC Overview
For more information on running ADCIRC and documentation, see the following links:
ADCIRC is available as a standalone app accessible via the DesignSafe front-end.
ADCIRC Inputs
An ADCIRC run is controlled by a variety of input files that can vary depending on the type of simulation being run. They all follow the naming convention fort.# where the # determines the type of input/output file. For a full list of input files for ADCIRC see the ADCIRC documentation. At a high level the inputs compose of the following:
- Base Mesh input files - Always present for a run. It will be assumed for the purpose of this UseCase that the user starts from a set of mesh input files.
- fort.14 - ADCIRC mesh file, defining the domain and bathymetry.
- fort.15 - ADCIRC control file, containing (most) control parameters for the run. This includes:
- Solver configurations such as time-step, and duration of simulation.
- Output configurations, including frequency of output, and nodal locations of output.
- Tidal forcing - At a minimum, ADCIRC is forced using tidal constituents.
- Additional control files (there are a lot more, just listing the most common here):
- fort.13 - Nodal attribute file
- fort.19, 20 - Additional boundary condition files.
- Meteorological forcing files - Wind, pressure, ice coverage, and other forcing data for ADCIRC that define a particular storm surge event.
- fort.22 - Met. forcing control file.
- fort.221, fort.222, fort.225, fort.22* - Wind, pressure, ice coverage (respective), and other forcing files.
The focus of this use case is to compile sets of storm surge events, each comprising different sets of forcing files, for a region of interest defined by a set of mesh control files.
PyADCIRC
The following use case uses the pyADCIRC python library to manage ADCIRC input files and get data from the data sources mentioned above. The library can be installed using pip:
$ pip install pyadcircThe pyadcirc.data contains functions to access two data sources in particular. First is NOAAs tidal gauge data for identifying storm surge. They provide a public API for accessing their data, for which pyADCIRC provides a python function and CLI (command line interface) wrapper around. The tidal signal at areas of interest over our domain will allow us to both identify potential storm surge events, and verify ADCIRC hind-casts with the real observations.
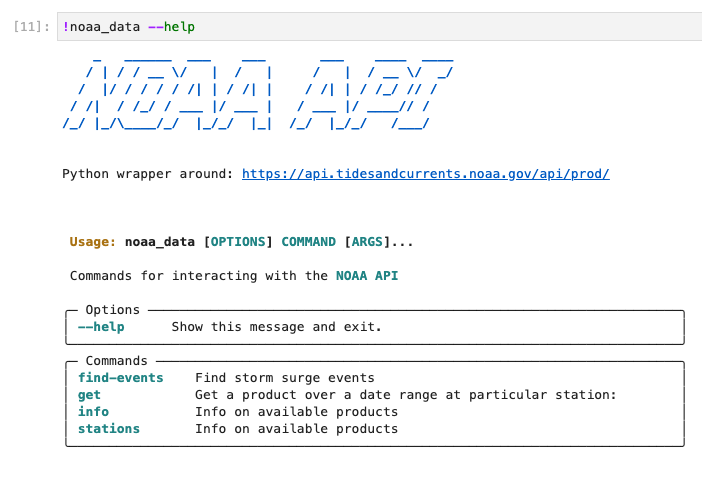
NOAA API CLI provided by the pyadcirc library. The noaa_data executable end point is created whenever pyadcirc is installed as library in an environment, providing a convenient CLI for interacting with the NOAA API that is well documented.
The second data source is NCAR’s CFSv1/v2 data sets for retrieving meteorological forcing files for identify storm surge events. An NCAR account is required for accessing this dataset. Make sure to go to NCAR's website to request an account for their data. You'll need your login information for pulling data from their repositories. Once your account is set-up, you'll want to store your credentials in a json file in the same directory as this notebook, with the name .ncar.json.
For example the file may look like:
{"email": "user@gmail.com", "pw": "pass12345"}Example Notebook: Creating ADCIRC DataSet
The example within this use case comprises of 4 main steps to create a data-set starting from a set of ADCIRC control input files. The notebook can be found at in the ADCIRC Use Case’s folder with the name Creating an ADCIRC DataSet.ipynb . Note that the notebook should be copied to the users ~/MyData directory before being able to use it (these steps are covered in the notebook).
The notebook covers the first two steps of this use case, namely identifying storm surge events and creating base input data sets to run using ADCIRC. We briefly overview the notebook’s results below.
Identifying storm surge events
The first stage of the notebook involves using the NOAA API wrapper provided by pyADCIRC to find storm surge events by looking at tidal gauge data in a region of interest. An example of an identified storm surge event, corresponding to Typhoon Merbok that hit the coast of Alaska in September 2022, is shown below.
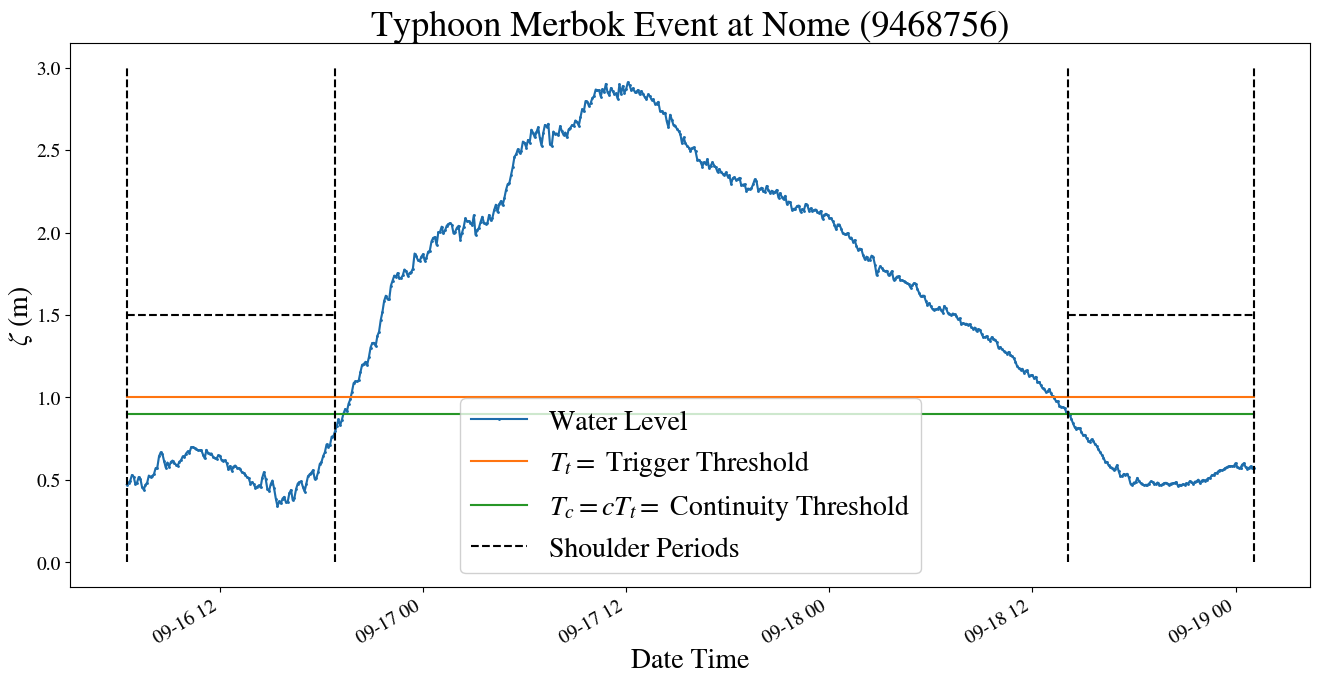
Result of identification algorithm for the range of dates containing Typhoon Merbok. The algorithm operates by defining a trigger threshold, along with other heuristics, by which to group distinct groups of storm surge events.
The algorithm presented is run on the storms that see the most frequent storm-surge activity over the coast of Alaska, Nome, Red Dog Dock, and Unalakleet. All events are compiled to give date ranges of storm surge events to produce ADCIRC hind-casts for.
Getting data forcing data
Having identified dates of interest, the notebook then uses the ncar library endpoint to pull meteorological forcing for the identified potential storm surge events. These are then merged with ADCIRC base input files (available at the published data set), to create input runs for an ensemble of ADCIRC simulations, as covered in the use case documentation on running ADCIRC ensembles in DesignSafe.
Organizing Data for publishing
Having a set of simulated ADCIRC hind-casts for one or more events, along with any additional analysis performed on the hind-cast data, the true power of DesignSafe as a platform can be realized by publishing your data. Publishing your data allows you and other researchers to reference its usage with a DOI. For ADCIRC, this is increasingly useful as more Machine Learning models are being built using ADCIRC simulation data.
This section will cover how to organize and publish an ADCIRC hind-cast dataset as created above. Note this dataset presented in this use case is a subset of the Alaska Storm Surge Data set that has been published, so please refrain from re-publishing data.
The steps for publishing ADCIRC data will be as follows
- Create a project directory in the DesignSafe data repository.
- Organize ADCIRC data and copy to project directory.
- Curate data by labeling and associating data appropriately.
While DesignSafe has a whole guide on how to curate and publish data, we note that the brief documentation below gives guidance on how to apply these curation guidelines to the particular case of ADCIRC simulation data.
Setting up Project Directory
First you’ll want to create a new project directory in the DesignSafe data repository.
Creating a new project in DesignSafe’s Data Depot.
Next we want to move ADCIRC inputs/outputs from your Jupyter instance where they were created into this project directory. We note that you must first restart your server if your moving data to a project directory that didn’t exist at the time from your server started, as that project directory won’t be in your ~/projects directory. Furthermore you’ll want to organize your folder structure in the command line before moving it to the project directory. See below for the recommended folder structure and associated data curation labels for publishing ADCIRC datasets.
.
├── Report.pdf -> Label as Report - PDF summarizing DataSet
├── mesh -> Label as Simulation Input (ADCIRC Mesh Type)
│ ├── fort.13
│ ├── fort.14
│ ├── fort.15
│ ├── fort.22
│ ├── fort.24
│ └── fort.25
├── inputs -> Label as Simulation Input (ADCIRC Meteorological Type)
│ ├── event000
│ │ ├── fort.15
│ │ ├── fort.221
│ │ ├── fort.222
│ │ └── fort.225
│ └── event001
│ ├── fort.15
│ ├── fort.221
│ ├── fort.222
│ └── fort.225
└── outputs -> Label as Simulation Output (ADCIRC Output)
├── event000
│ ├── fort.61.nc
│ ├── ...
│ ├── maxele.63.nc
│ ├── maxrs.63.nc
│ ├── maxvel.63.nc
│ ├── maxwvel.63.nc
│ └── minpr.63.nc
└── event001
├── fort.61.nc
├── ...
├── maxele.63.nc
├── maxrs.63.nc
├── maxvel.63.nc
├── maxwvel.63.nc
└── minpr.63.nc
└── Analysis -> Label as Analysis any notebooks/code/images.
├── OverviewNotebook.ipynb - Analysis over all events.
├── event000
│ ├── ExampleNotebook.ipynb - Event specific analysis.
│ ├── ...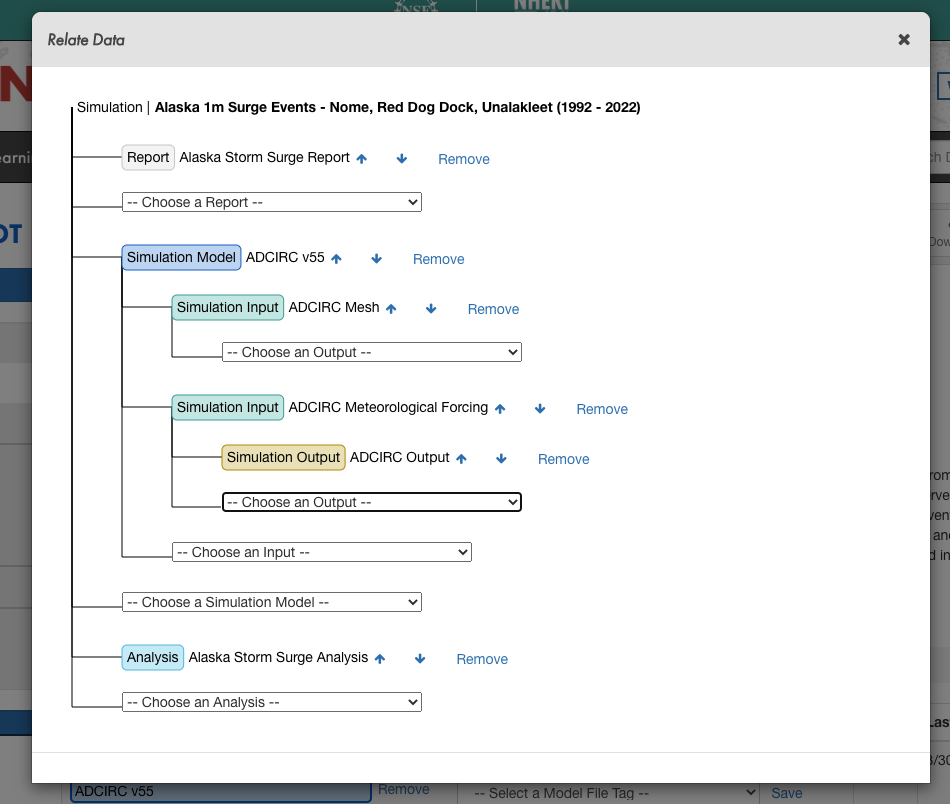
Example data relation diagram for an ADCIRC Simulation DataSet
Visualizing Surge for Regional Risks
Visualizing Surge for Regional Risks
Integration of QGIS and Python Scripts to Model and Visualize Storm Impacts on Distributed Infrastructure Systems
Catalina González-Dueñas and Jamie E. Padgett - Rice University
Miku Fukatsu - Tokyo University of Science
This use case study shows how to automate the extraction of storm intensity parameters at the structure level to support regional risk assessment studies. This example leverages QGIS and python scripts to obtain the surge elevation and significant wave height from multiple storms at specific building locations. The case study also shows how to visualize the outputs in QGIS and export them as a web map.
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
| Scope | Notebook |
|---|---|
| Read Surge | Surge_Galv.ipynb |
| Read Wave | Wave_Galv.ipynb |
DesignSafe Resources
The following DesignSafe resources are leveraged in this example:
Geospatial data analysis and Visualization on DS - QGIS
Jupyter notebooks on DS Jupyterhub
Background
Citation and Licensing
-
Please cite González-Dueñas and Padgett (2022) to acknowledge the use of any resources from this use case.
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License.
Description
This case study aims to support pre-data processing workflows for machine learning applications and regional risk analysis. When developing predictive or surrogate models for the response of distributed infrastructure and structural systems, intensity measures (IMs) need to be associated with each component of the system (e.g., buildings, bridges, roads) under varying hazard intensity or different hazard scenarios. To accomplish this and given the different resolutions of the hazard and infrastructure data, geographical tools need to be used to associate the intensity measures with the distributed infrastructure or portfolio components. In this case study, python codes were developed to automate geospatial analysis and visualization tasks using QGIS.
This case study is divided into four basic components:
- Introduction and workflow of analysis
- Storm data analysis using Jupyter notebooks
- Geospatial analysis via QGIS
- Visualization of the outputs
Introduction and workflow of analysis
In this example, the automated procedure to extract intensity measures is leveraged to obtain the maximum surge elevation and significant wave height at specific house locations for different storm scenarios. The surge elevation and the significant wave height are important parameters when evaluating the structural performance of houses under hurricane loads, and have been used to formulate different building fragility functions (e.g., Tomiczek, Kennedy, and Rogers (2014); Nofal et al. (2021)){target=_blank}. As a proof of concept, the intensity measures (i.e., surge elevation and significant wave height) will be extracted for 3 different storms using the building portfolio of Galveston Island, Texas. The storms correspond to synthetic variations of storm FEMA 33, a probabilistic storm approximately equivalent to a 100-year return period storm in the Houston-Galveston region. The storms are simulated using ADCIC+SWAN numerical models of storm FEMA33, with varying forward storm velocity and sea-level rise. For more details on the storm definition, the user can refer to Ebad et al. (2020) and González-Dueñas and Padgett (2021).
In order to relate the storm data to the building portfolio data, it is necessary to convert the storm outputs to a surface data and then extract at the locations of interest. First, the output files from the ADCIRC+SWAN simulation corresponding to the surge elevation (file fort.63.nc) and significant wave height (file swan_HS.63.nc) need to be converted to a format that can be exported to a GIS (Geographical Information System) software. This pre-processing of the storm data provides the surge elevation and significant wave height in each of the grid points used to define the computational domain of the simulation in a vector data format. Since these points (i.e., ADCIRC+SWAN grid) have a different spatial resolution than the infrastructure system under analysis (i.e., building locations), the storm outputs are converted to a surface data format and then the value at each building location is extracted from it. This is repeated for each one of the storms under analysis and then the ouput data (IMs at each building location) is exported as a csv file. This file is used to support further analysis in the context of risk assessment or machine learning applications, as predictors or response of a system. The workflow of analysis is as follows:
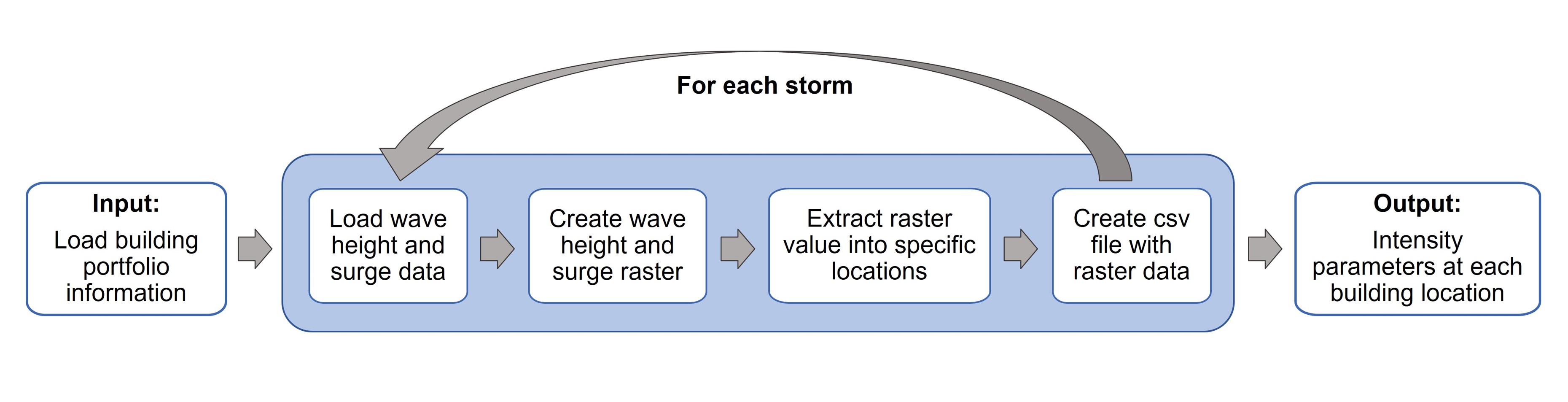
Storm data analysis using Jupyter notebooks
To read the ADCIRC+SWAN storm simulation outputs, two Jupyter notebooks are provided, which can extract the maximum surge elevation and significant wave height values within a particular region. The Read_Surge Jupyter notebook takes as an input the fort.63.nc ADCIRC+SWAN output file and provides a csv file with the maximum surge elevation value at each of the points within the region specified by the user. Specifying a region helps to reduce the computational time and to provide the outputs only on the region of interest for the user. Similarly, the Read_WaveHS Jupyter notebook, reads the swan_HS.63.nc file and provides the maximum significant wave height in the grid points of the specified area. Direct links to the Notebooks are provided above.
To use the Jupyter notebooks, the user must:
- Create a new folder in My data and copy the Jupyter notebooks from the Community Data folder
- Ensure that the fort.63.nc and swan_HS.63.nc are located in the same folder as the Jupyer notebooks
- Change the coordinates of the area of interest in [6]:
### Example of a polygon that contains Galveston Island, TX (The coordinates can be obtained from Google maps)
polygon = Polygon([(-95.20669, 29.12035), (-95.14008, 29.04294), (-94.67968, 29.35433), (-94.75487, 29.41924), (-95.20669, 29.12035)])### In this example, the output name of the csv surge elevation file is "surge_max"
with open('surge_max.csv','w') as f1:Geospatial analysis via QGIS
Opening a QGIS session in DesignSafe
To access QGIS via DesignSafe go to Workspace -> Tools & Applications -> Visualization -> QGIS Desktop 3.16. You will be prompted the following window:
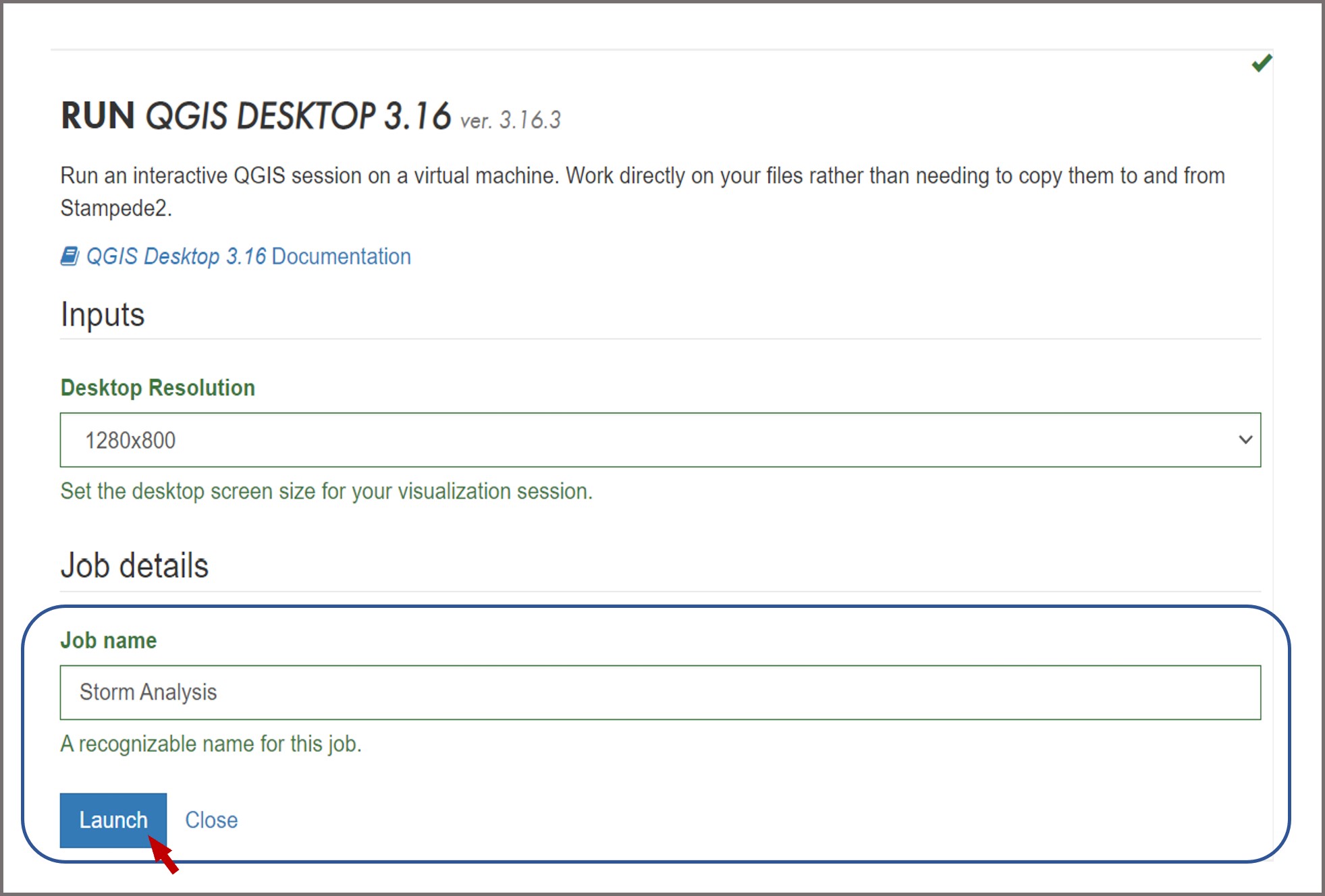
Change the desktop resolution according to your screen size preferences, provide a name for your job, and hit Launch when you finish. After a couple of minutes your interactive session will start, click Connect:
You will be directed to an interactive QGIS session, create a new project by clicking the New Project icon or press Ctrl+N:
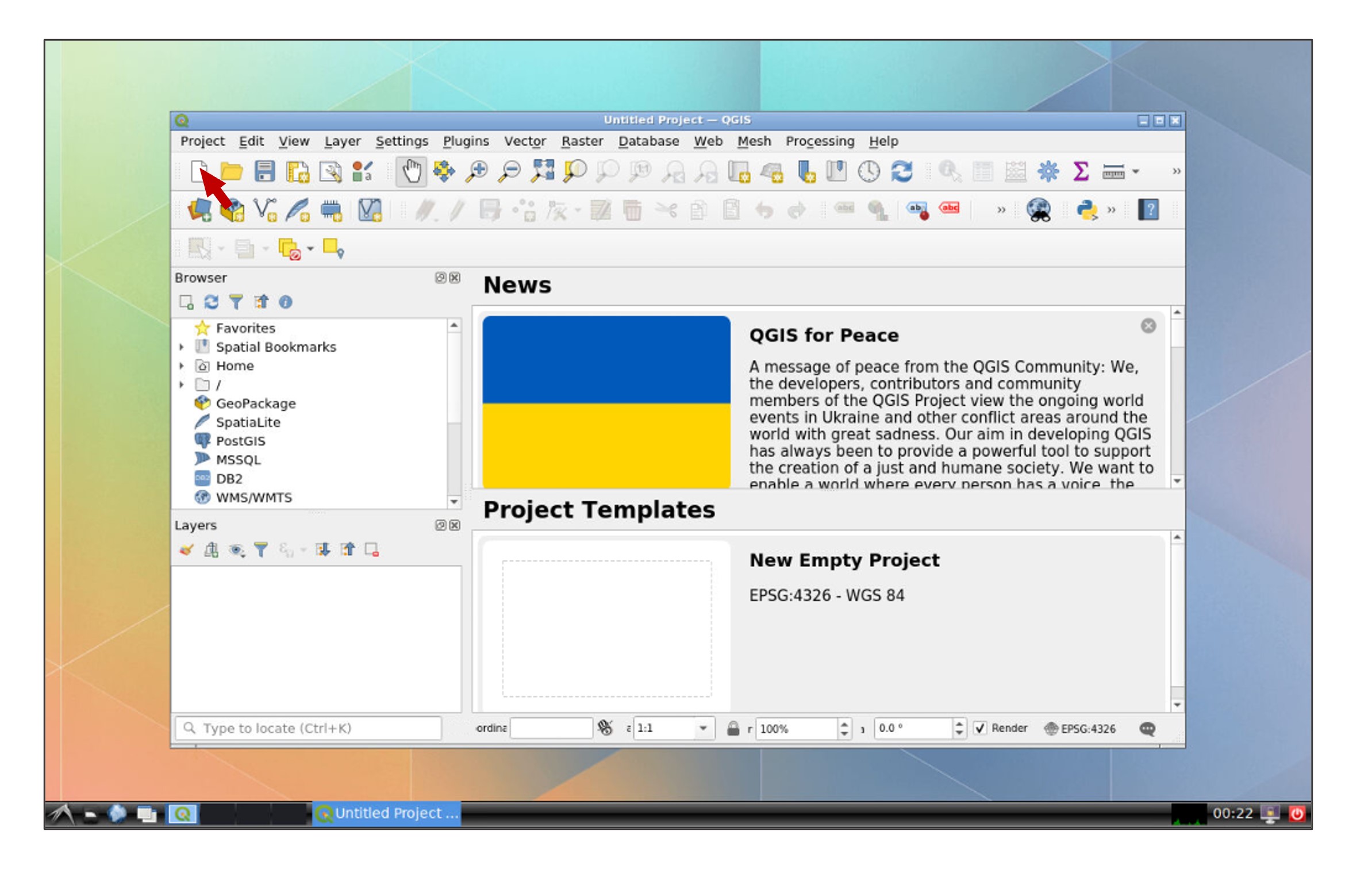
Modify user inputs and run the python script
A python script called IM_Extract is provided to extract the desired IMs at specific locations. Follow these steps to use this code:
- Create a folder to store the outputs of the analysis in your My data folder in DS
- Provide a csv file that specifies the points for which you wish to obtain the intensity measures. This file should be in the following format (see the Complete_Building_Data file for an example of the building stock of Galveston Island, TX): a. The first column should contain an ID (e.g., number of the row) b. The second column corresponds to the longitude of each location c. The third column corresponds to the latitude of each location
- Create a folder named Storms in which you will store the data fromt the different storms
- Within the Storms folder, create a folder for each one of the storms you wish to analyze. Each folder should contain the output csv files from the ADCIRC+SWAN simulations (e.g., surge_max.csv, wave_H_max.csv). In our case study, we will use three different storms.
Once the folder of analysis is created in your Data Depot, we can proceed to perform the geospatial analysis in QGIS. Open the python console within QGIS, click the Show Editor button, and then click Open Script:
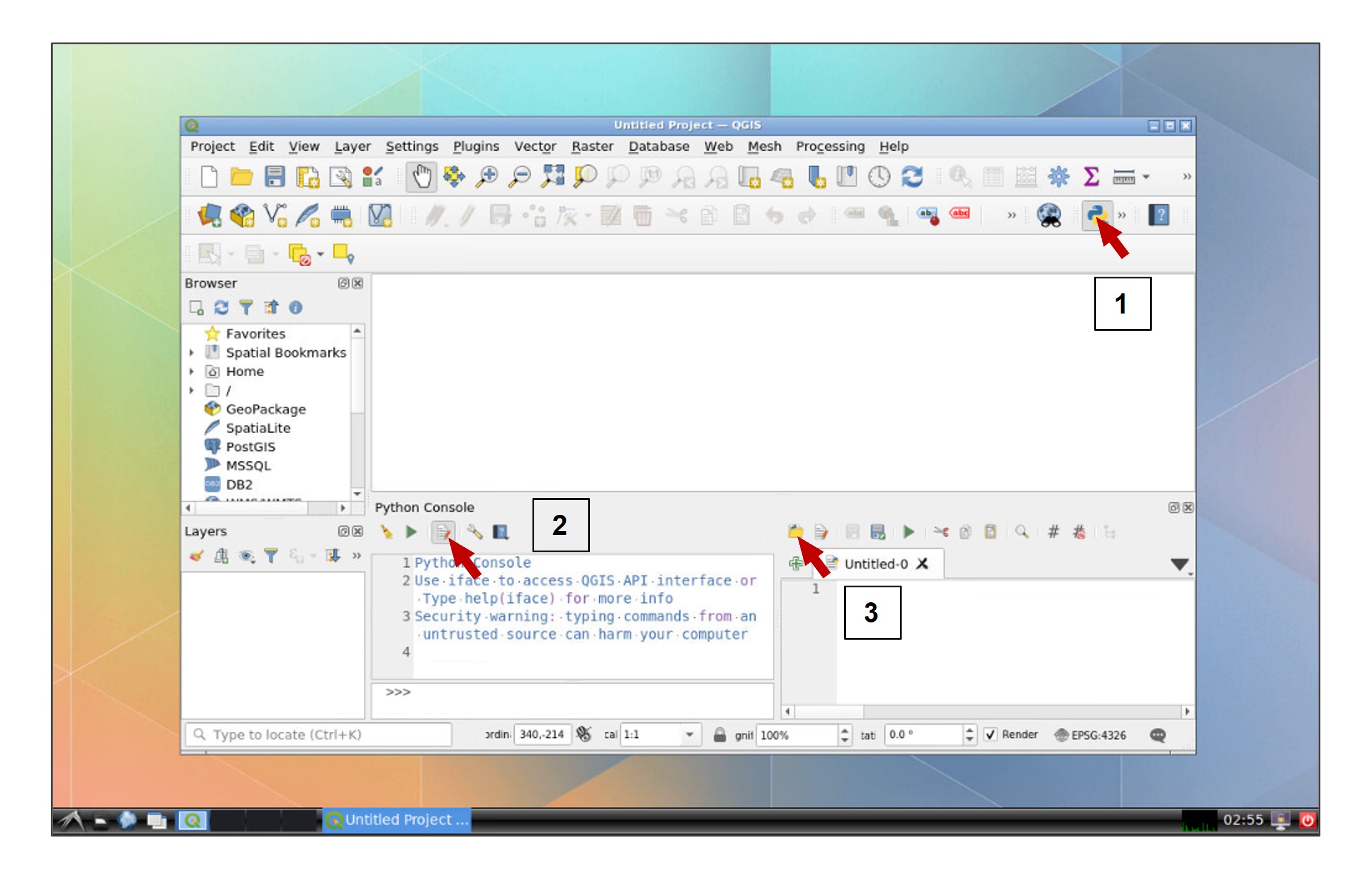
In the file explorer, go to your data folder and open the IM_Extract script:
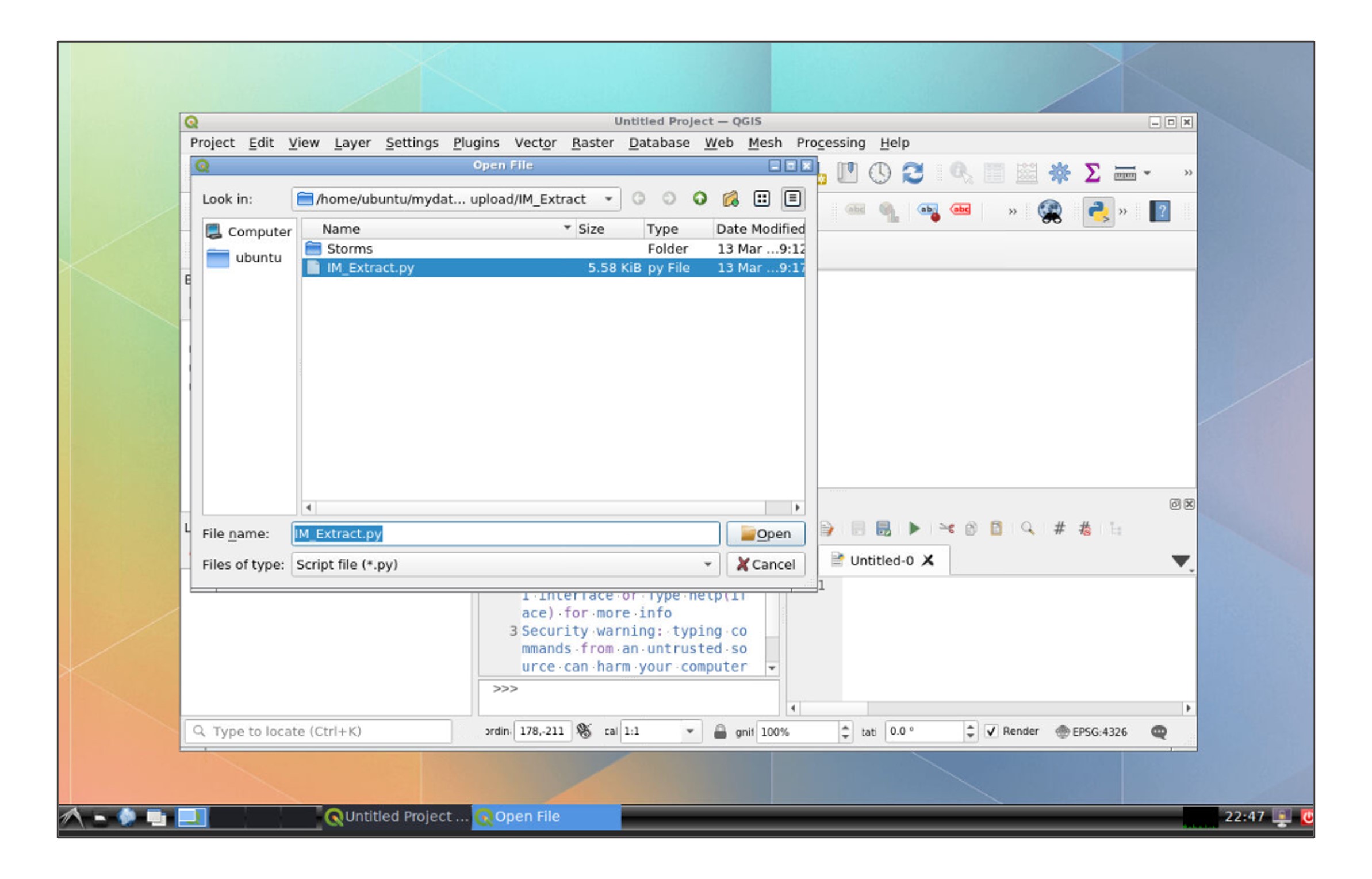
Modify the path of the folder for your own data folder in line 17:
path= r"/home/ubuntu/mydata/**name of your folder**"### Line 22
# 2. Change cell size (defalt is 0.001)
cell_size = 0.001
### Line 75
# Interpolation method
alg = "qgis:tininterpolation"Once you finish the modifications, click Run Script.
Visualization of the outputs
Once the script finish running, the time taken to run the script will appear in the python console and the layers created in the analysis will be displayed in the Layers section (left-bottom window) in QGIS:
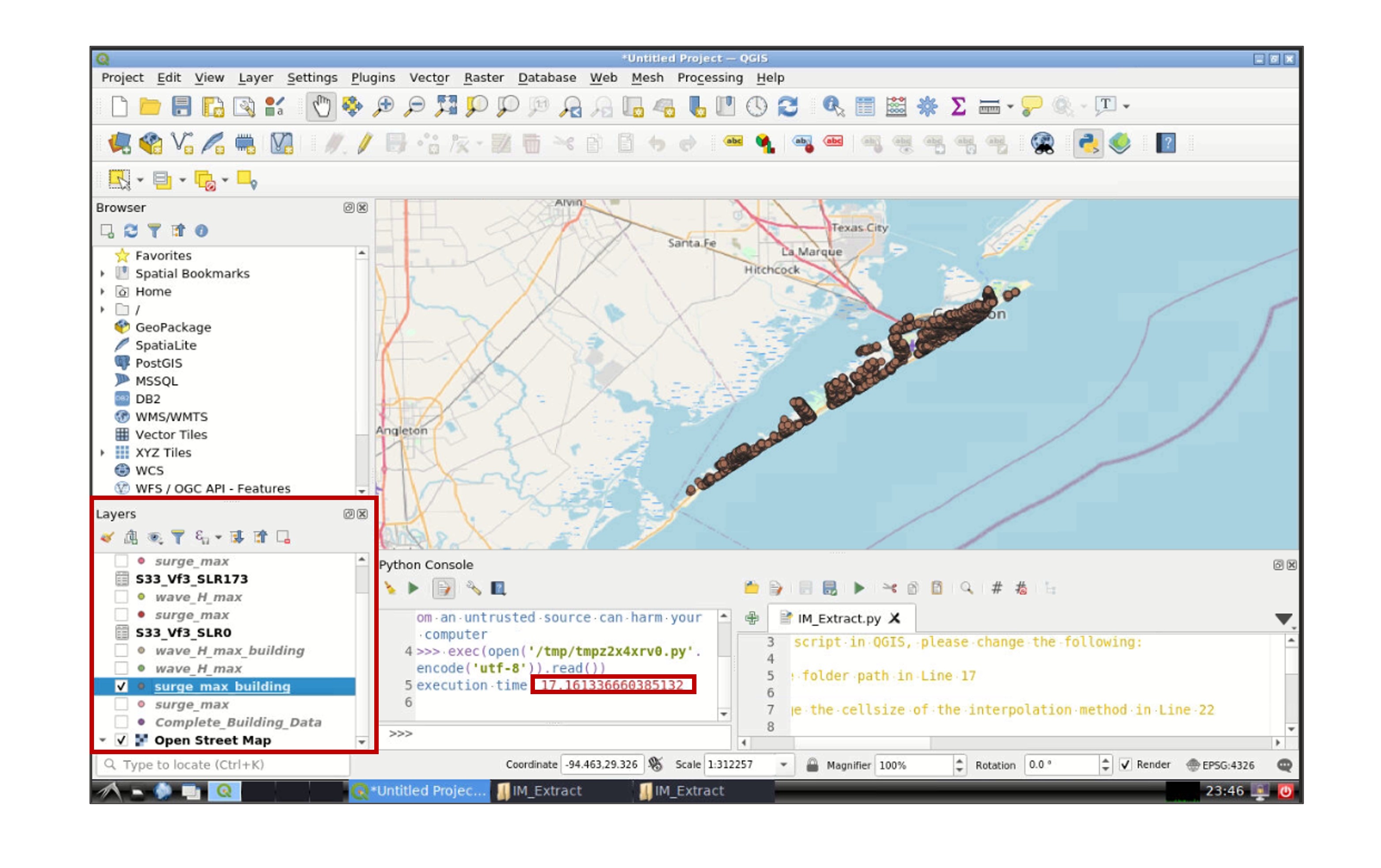
Right click one of the layers, and go to Properties -> Symbology to modify the appearance of the layer (e.g., color, size of the symbol):
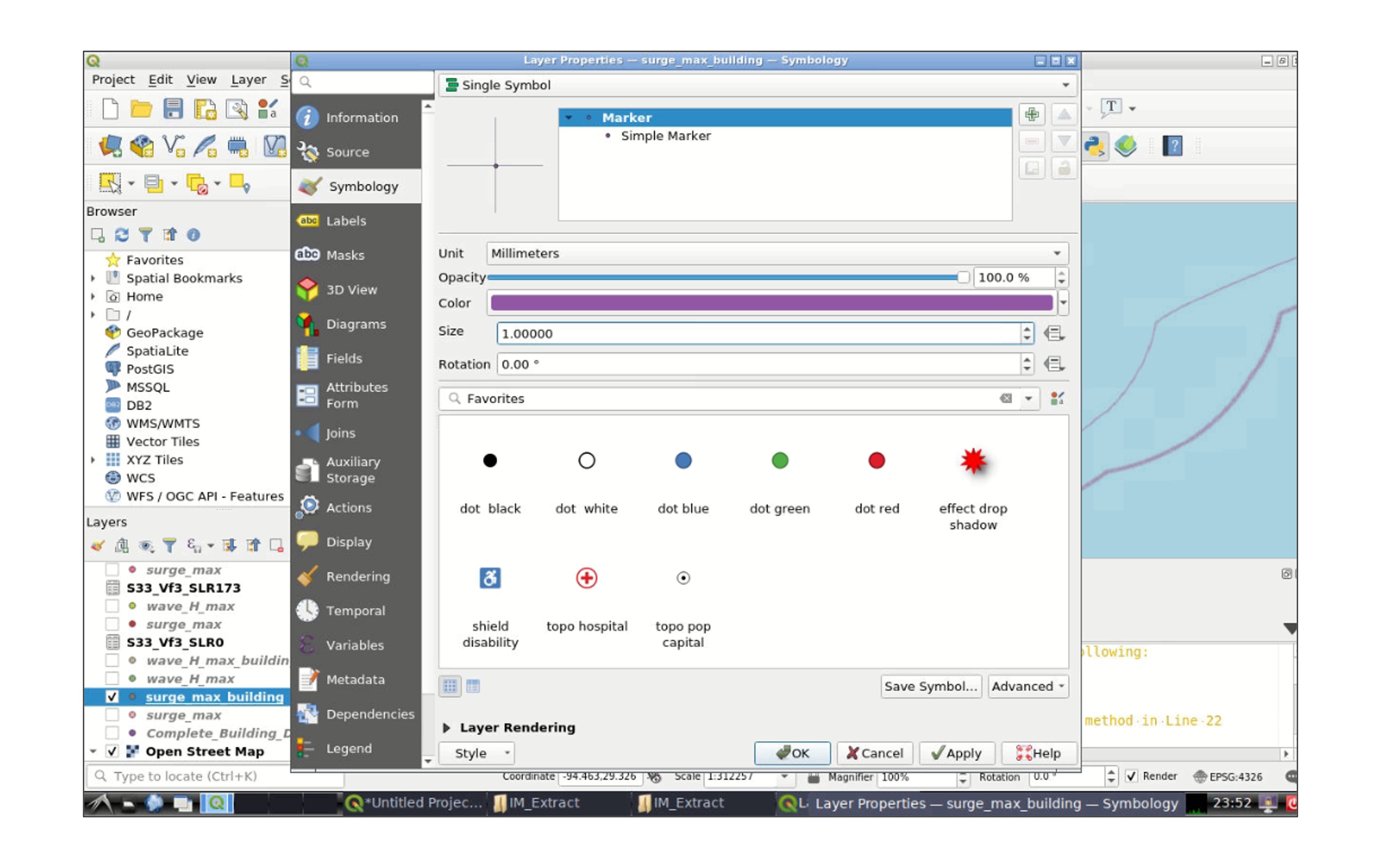
Click OK when you finish the modifications. You will be directed to the main window again, go to the the toolbar and click Plugins -> Manage and Install Plugins. In the search tab type qgis2web, select the plugin, and click Install Plugin:
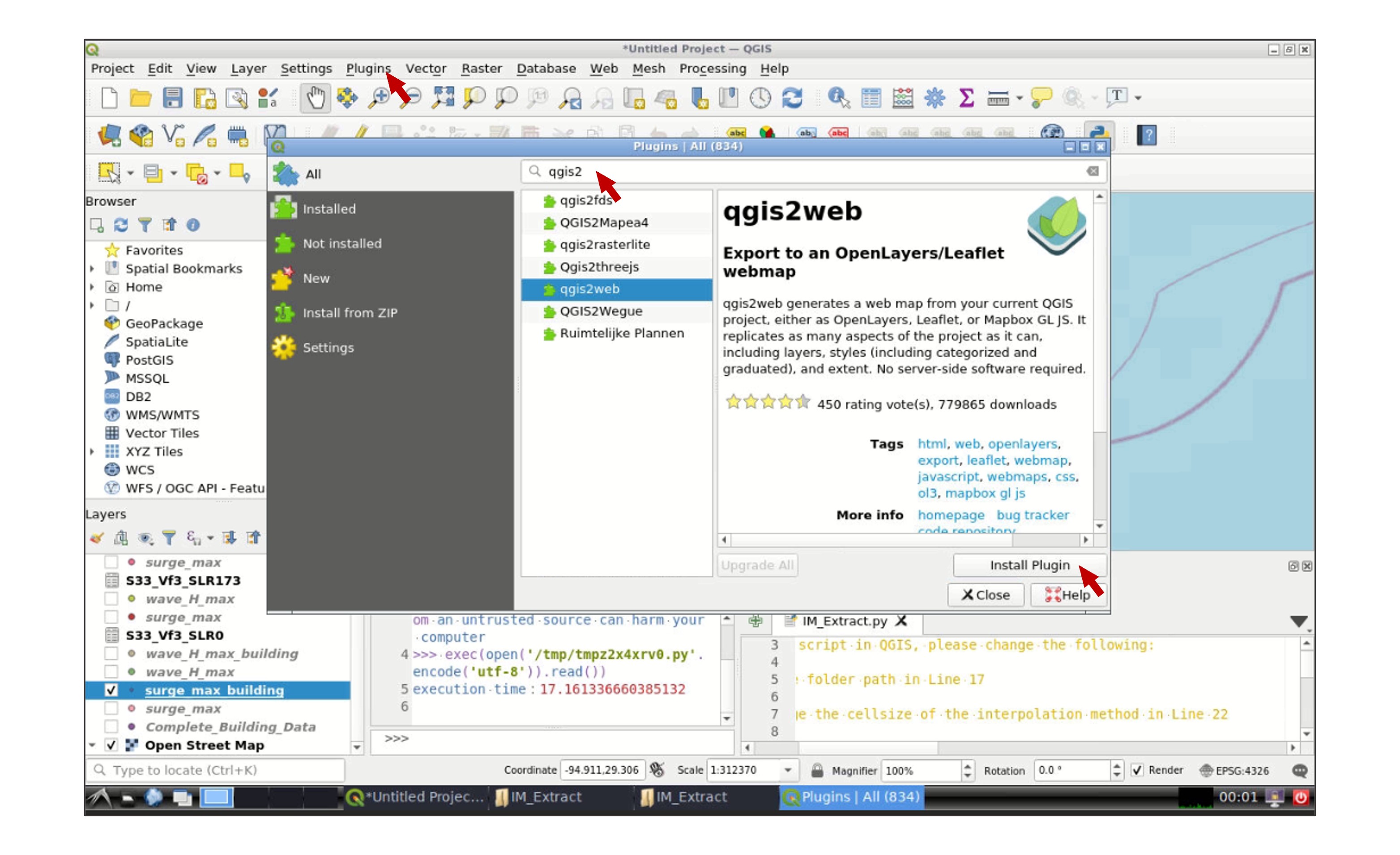
Go to Web -> qgis2web -> Create web map:
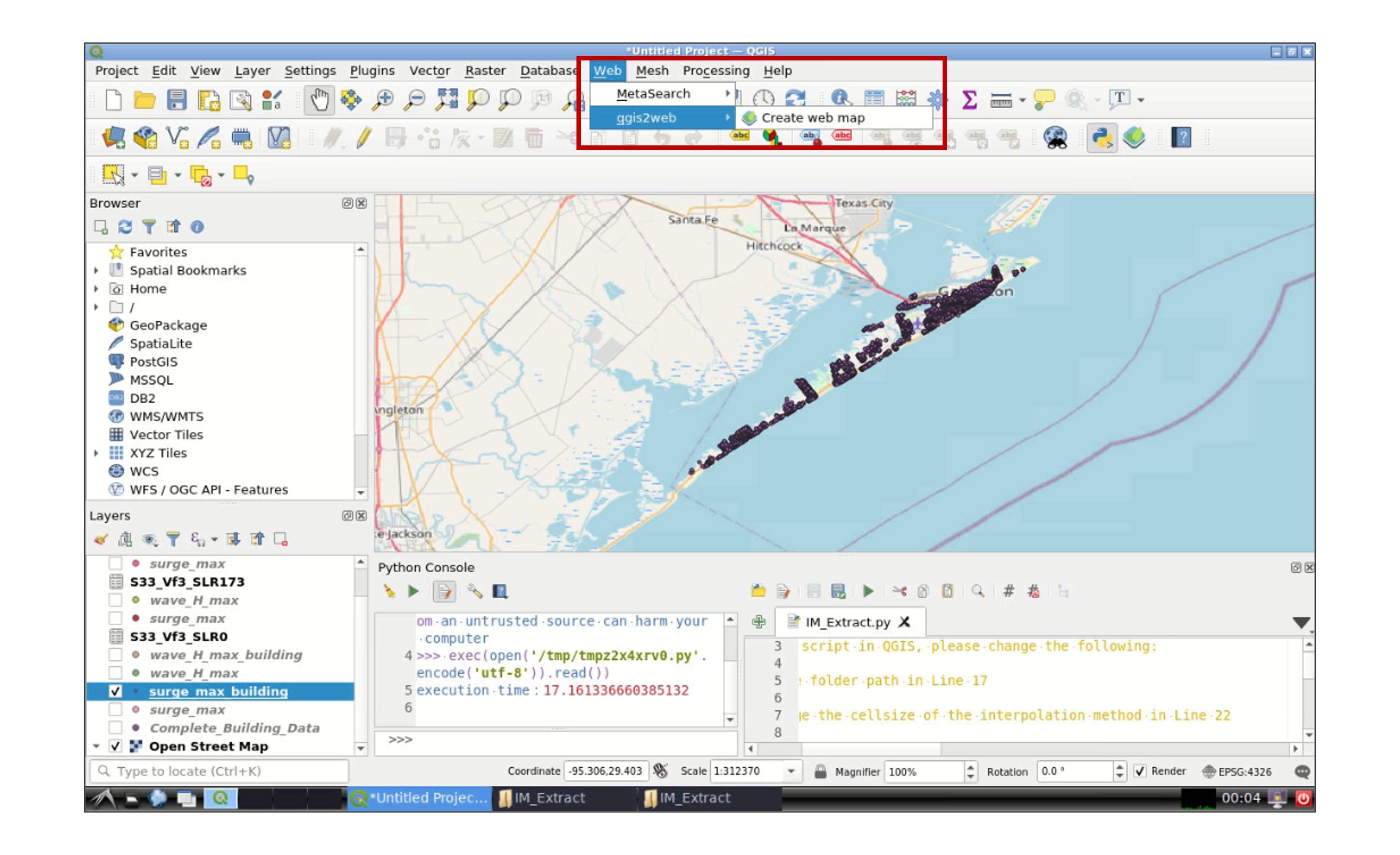
In the new window, select the layer(s) that you wish to export in the Layers and Groups tab, and modify the appearance of the map in the Appearance tab. Then go to the Export tab and click in the icon next to Export to folder and select your working data folder:
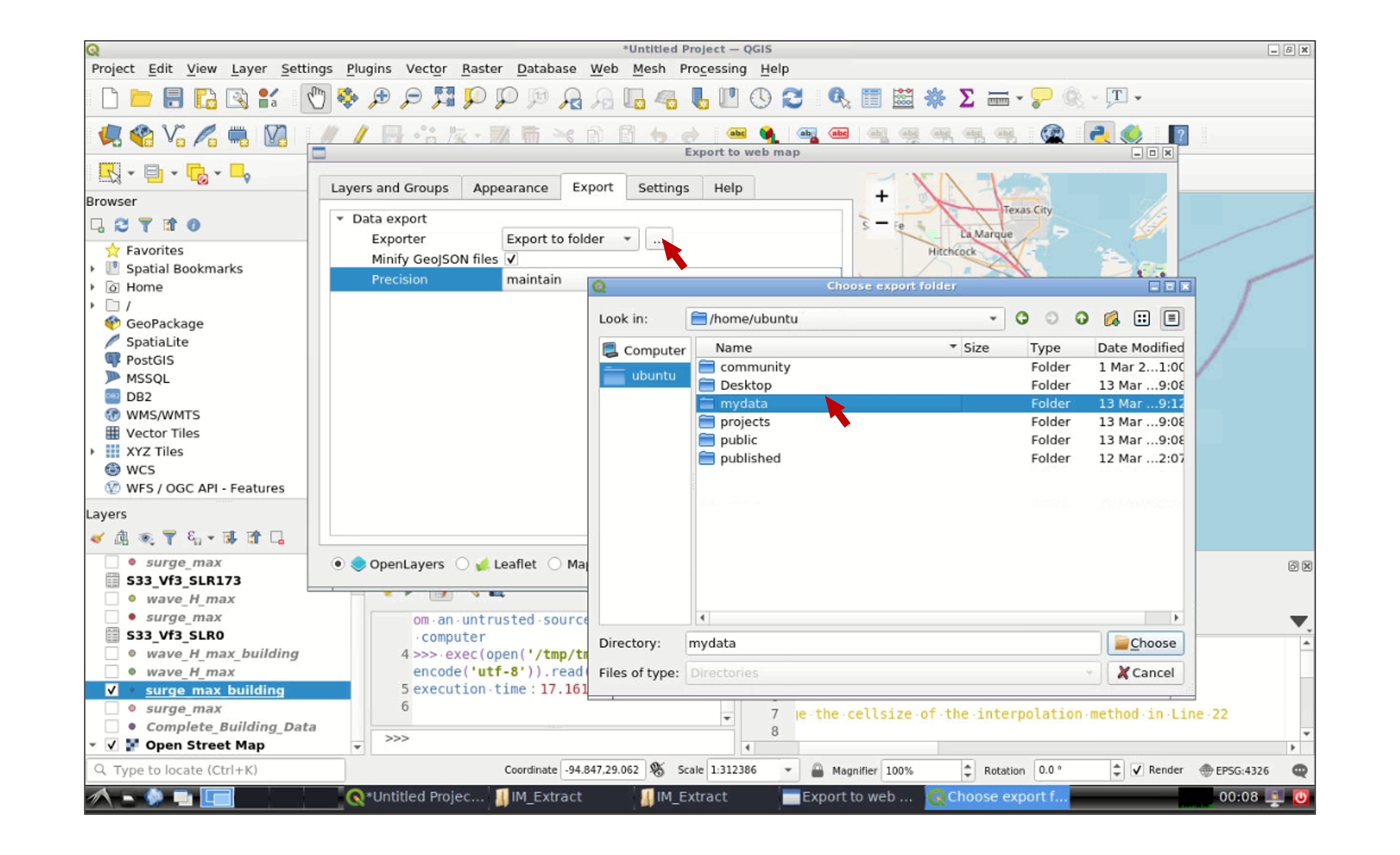
Once you finish, a new web explorer window will open in your interactive session with the exported QGIS map:
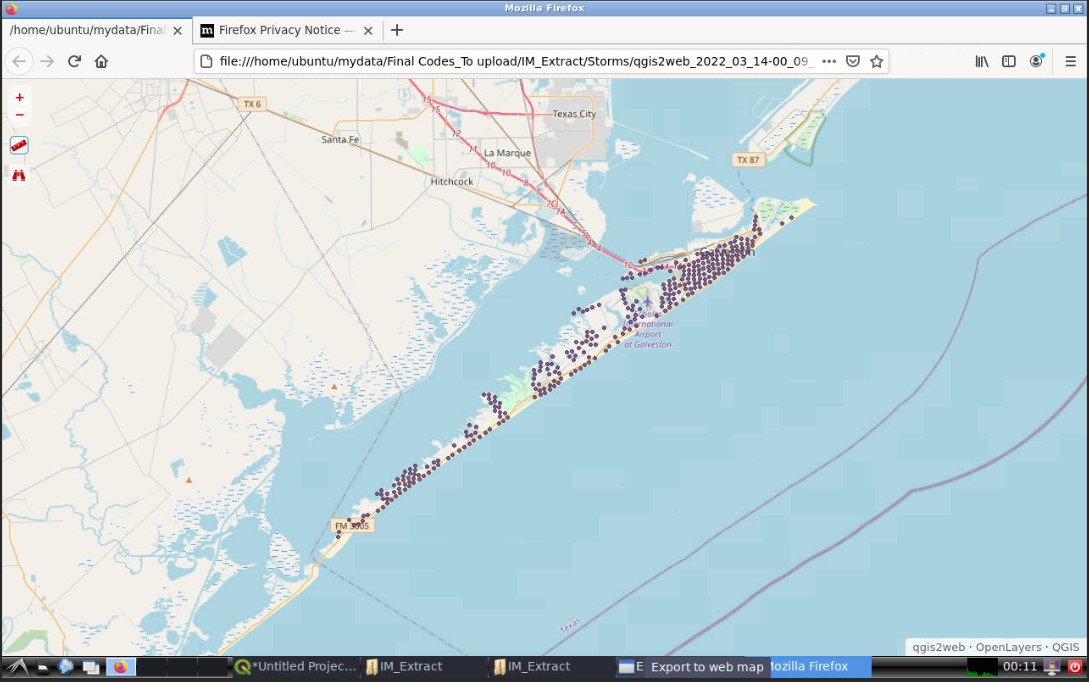
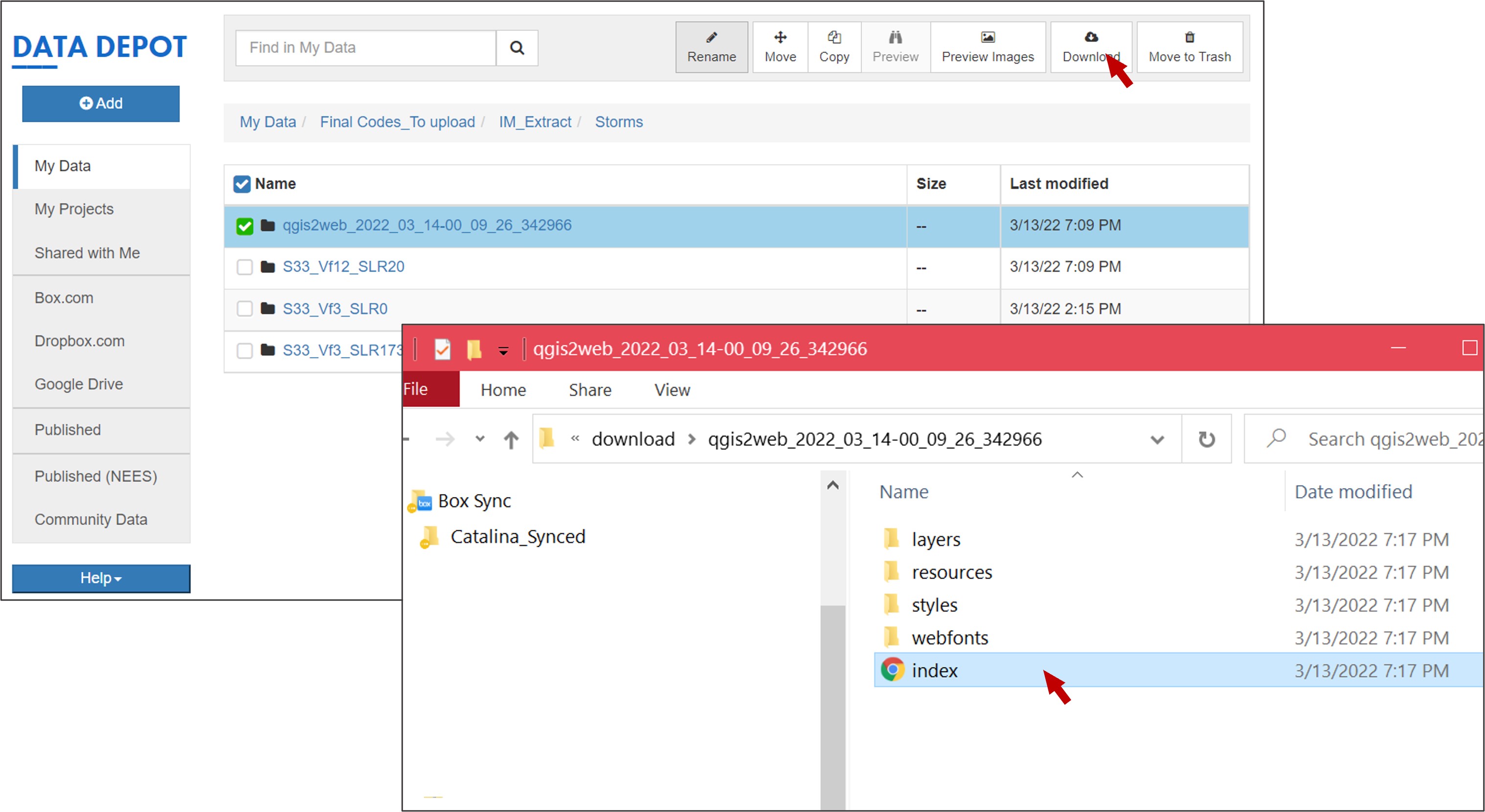
Go to your working folder in the Data Depot, a new folder containing the web map will be created. You can download the folder and double click Index, the web map you created will be displayed in the web explorer of your local computer:
CFD Analysis of Winds on Structures
CFD Analysis of Winds on Structures
CFD Simulations using the Jupyter Notebooks
Fei Ding - NatHaz Modeling Laboratory, University of Notre Dame
Ahsan Kareem - NatHaz Modeling Laboratory, University of Notre Dame
Dae Kun Kwon - NatHaz Modeling Laboratory, University of Notre Dame
OpenFOAM is the free, open source CFD software and is popularly used for computationally establishing wind effects on structures. To help beginners overcome the challenges of the steep learning curve posed by OpenFOAM and provide users with the capabilities of generating repetitive jobs and advanced functions, this use case example presents the work to script the workflow for CFD simulations using OpenFOAM in the Jupyter Notebooks. The developed two Jupyter Notebooks can aid in determining inflow conditions, creating mesh files for parameterized building geometries, and running the selected solvers. They can also contribute to the education for CFD learning as online resources, which will be implemented in the DesignSafe.
All files discussed in this use case are shared at Data Depot > Community Data. It is recommended that users make a copy of the contents to their directory (My Data) for tests and simulations.
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
| Scope | Notebook |
|---|---|
| Jupyter PyFoam Example | Jupyter_PyFoam.ipynb |
| Use case Example | OpenFOAM_Run_example.ipynb |
DesignSafe Resources
The following DesignSafe resources were leveraged in developing this use case.
OpenFoam
ParaView
Jupyter notebooks on DS Juypterhub
Background
Citation and Licensing
- Please cite Ding and Kareem (2021) to acknowledge the use of any resources from this use case.
- Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
- This software is distributed under the GNU General Public License.
OpenFOAM with the Jupyter Notebook for creating input environments
The overall concept of the OpenFOAM workflow may be expressed as physical modeling-discretisation-numerics-solution-visualization as shown in Fig. 1.
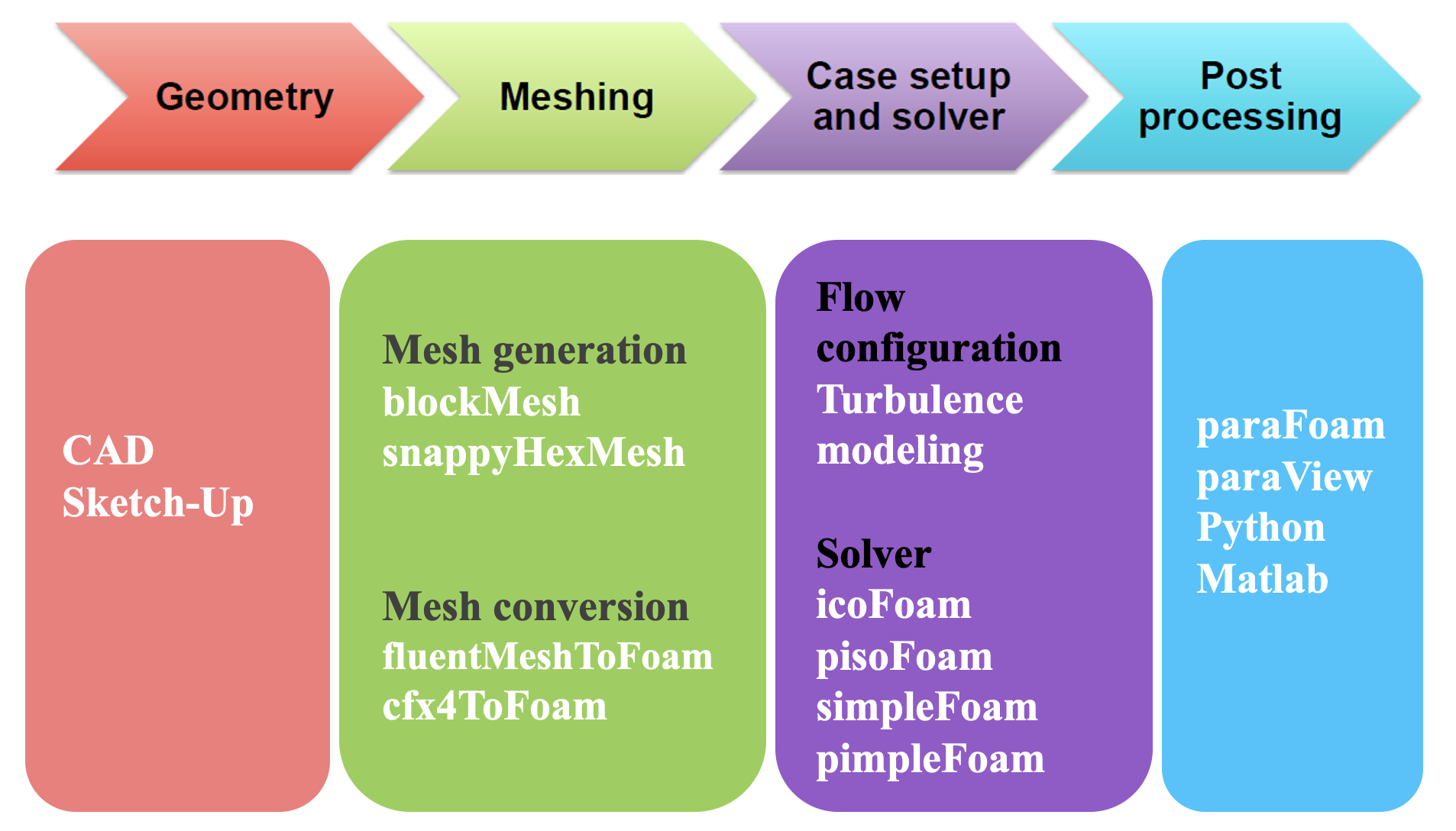
Fig. 1 OpenFOAM workflow for CFD modeling
Prerequisite to run OpenFOAM simulation
To run a CFD simulation using OpenFOAM, three directories (and associated input files) named 0, constant and system should be predefined by users. If the root directory of the directories is DH1_run, then it has the following directory structure [1].
DH1_run # a root directory
- 0 # initial and boundary conditions for CFD simulations
- constant # physical properties and turbulence modeling
- system # run-time control (parallel decomposition) and solverpisoFoam which is a transient solver for incompressible and turbulent flows and simpleFoam as a steady-state solver. Parallel computations in OpenFOAM allow the simulation to run in the distributed processors simultaneously.
Introducing advanced utilities to CFD modeling using PyFoam
Jupyter Notebooks can provide an interpretable and interactive computing environment to run a CFD simulation using the OpenFOAM. To introduce such flexibilities and bring maximum automation to CFD modeling using the OpenFOAM, an OpenFOAM library named PyFoam [2] can be used in the Jupyter Notebooks, which can introduce advanced tools for CFD modeling. With the aid of the PyFoam, the goal is to achieve an end-to-end simulation in which the Jupyter Notebooks can manipulate dictionaries in OpenFOAM based on the user's input as regular Python dictionaries without looking into the OpenFOAM C++ libraries (Fig. 2).
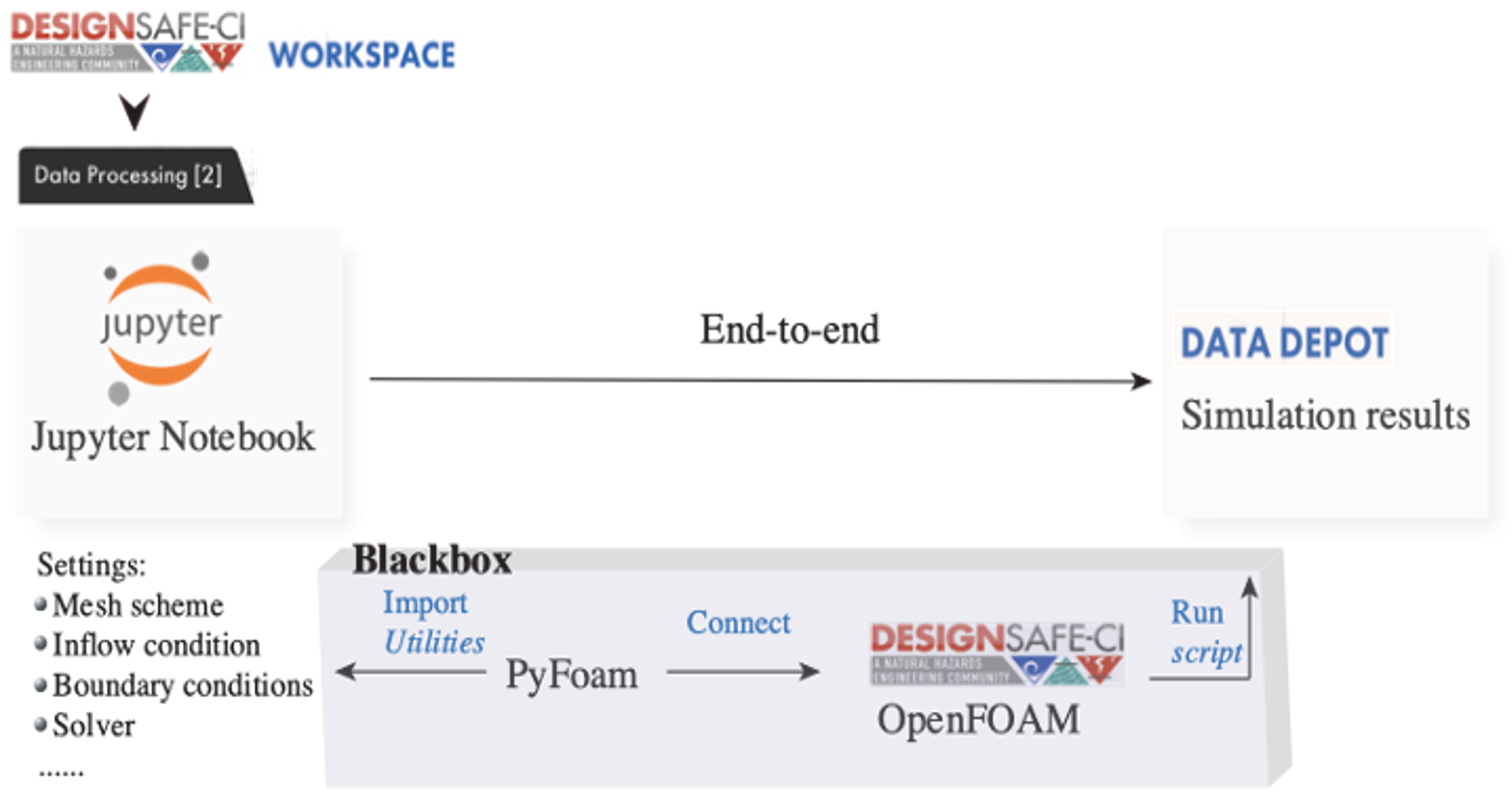
Fig. 2 Schematic of an end-to-end flow simulation implemented in the Jupyter Notebooks
Jupyter Notebook example for advanced utilities
For better understanding, A Jupyter Notebook example, Jupyter_PyFoam.ipynb, is provided that facilitates the automated CFD modeling with the aid of advanced utilities. Automated mesh generation and inflow configuration in the Jupyter Notebooks are explored through the case study of a rectangular building's cross-section.
In addition, a baseline model housed in DH_Baseline directory is provided that can be used to generate an input environment for an OpenFOAM simulation.
It is worth noting that DesignSafe recently introduced a Jupyterhub Spawner for users to run one of two Jupyter server images. To run Jupyter Notebooks for CFD presented in this document, users should use the Classic Jupyter Image as the Jupyter server.
Using PyFoam utilities in the Jupyter Notebook
At first, PyFoam and other modules should be imported into a Notebook, e.g.:
import sys
sys.path.append('/home1/apps/DesignSafe')
import PyFoam
import os, shutil, scipy.io, math, glob
import numpy as np
from pylab import *
from PyFoam.Execution.UtilityRunner import UtilityRunner
from PyFoam.Execution.BasicRunner import BasicRunner
from PyFoam.RunDictionary.SolutionDirectory import SolutionDirectory
from PyFoam.RunDictionary.SolutionFile import SolutionFile
from PyFoam.RunDictionary.BlockMesh import BlockMesh
from os import path
from subprocess import Popen
from subprocess import callMesh generation
To allow users to edit the dimensions of the rectangular building's cross-section, m4-scripting is employed for parameterization in OpenFOAM. To achieve it, case directories of the baseline geometry which is a square cross-section were first copied to the newly created case directories. The controlling points for mesh topology are functions of the input geometric variables. M4-scripting then manipulates the blockMeshDict dictionary, from which values of the controlling points were assigned as shown in Fig. 3.
In the end, the blockMeshDict dictionary file, which is the file for specifying the mesh parameter and used to generate a mesh in OpenFOAM, is built by executing m4-script command, e.g.,
cmd='m4 -P blockMeshDict.m4 > blockMeshDict'
pipefile = open('output', 'w')
retcode = call(cmd,shell=True,stdout=pipefile)
pipefile.close()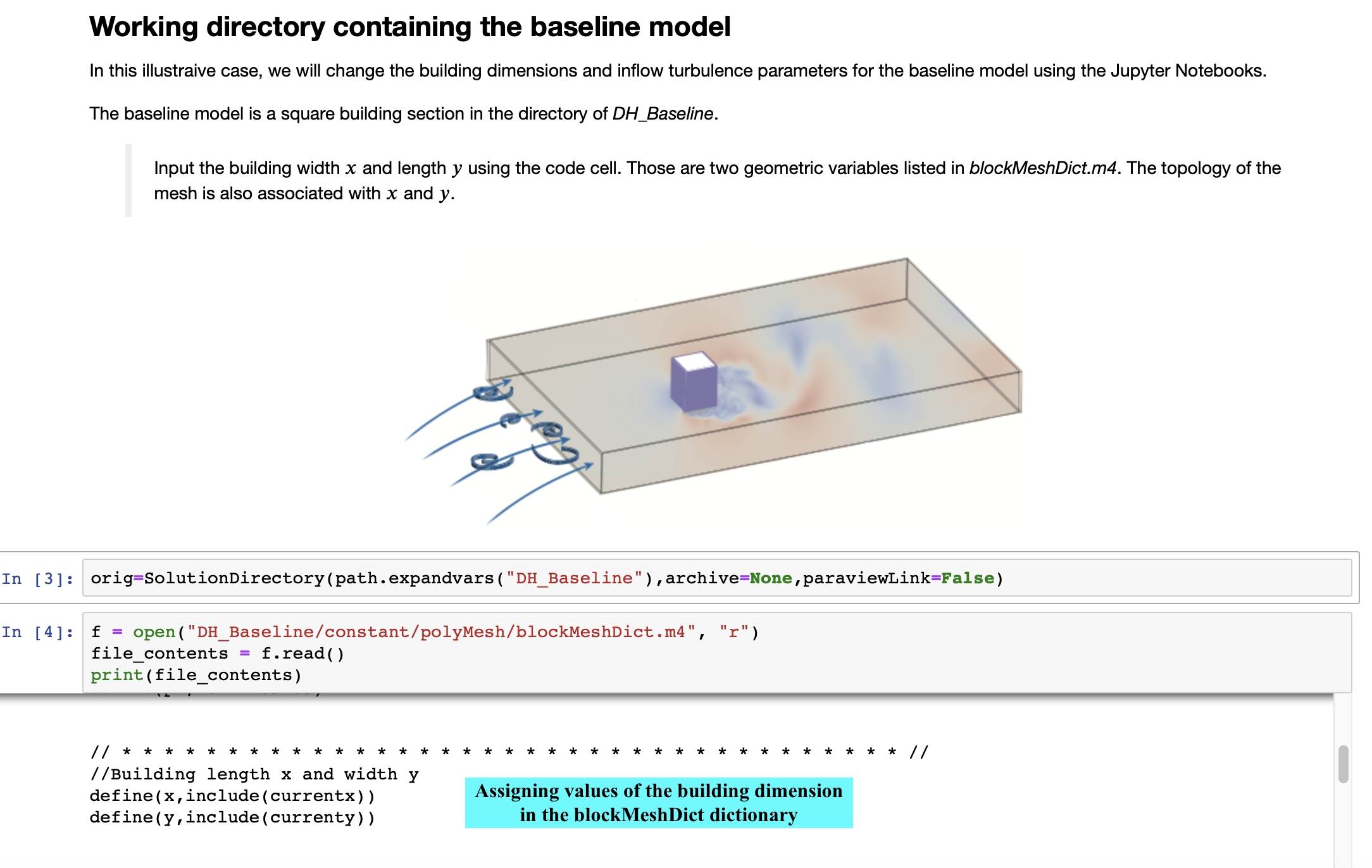
Fig. 3 Use of m4-scripting for automated mesh generation
Setup inflow condition
To edit the inflow turbulence properties based on the user's input, PyFoam is employed to set the inflow boundary conditions. In this example, k-ω SST model is selected for turbulence modeling for a demonstration, hence the two inflow turbulence parameters k and ω are modified at different inflow conditions by calling the replaceBoundary [2] utility in the PyFoam. Part of the codes is shown in the following:
from PyFoam.Execution.UtilityRunner import UtilityRunner
from PyFoam.Execution.BasicRunner import BasicRunner
from PyFoam.RunDictionary.SolutionDirectory import SolutionDirectory
from PyFoam.RunDictionary.SolutionFile import SolutionFile
### change the values k and ω at the inlet
os.chdir(case)
dire=SolutionDirectory(case,archive=None)
sol=SolutionFile(dire.initialDir(),”k”)
sol.replaceBoundary(”inlet”,”%f” %(k))
sol=SolutionFile(dire.initialDir(),”omega”)
sol.replaceBoundary(”inlet”,”%f” %(omega))More detailed information can be found in the Jupyter_PyFoam.ipynb. The Notebook also illustrates how to prepare multiple models to simulate simultaneously and their automatic generation of input environments.
Use case of an OpenFOAM simulation with Jupyter Notebook in the DesignSafe workspace
Description
A use case example is a URANS simulation for wind flow around a rectangular building's cross-section, which is implemented at a Jupyter Notebook, OpenFOAM_Run_example.ipynb. The input environments are prepared at DH1_run directory. The test rectangular cross-section model and its mesh are shown in Fig. 4. |
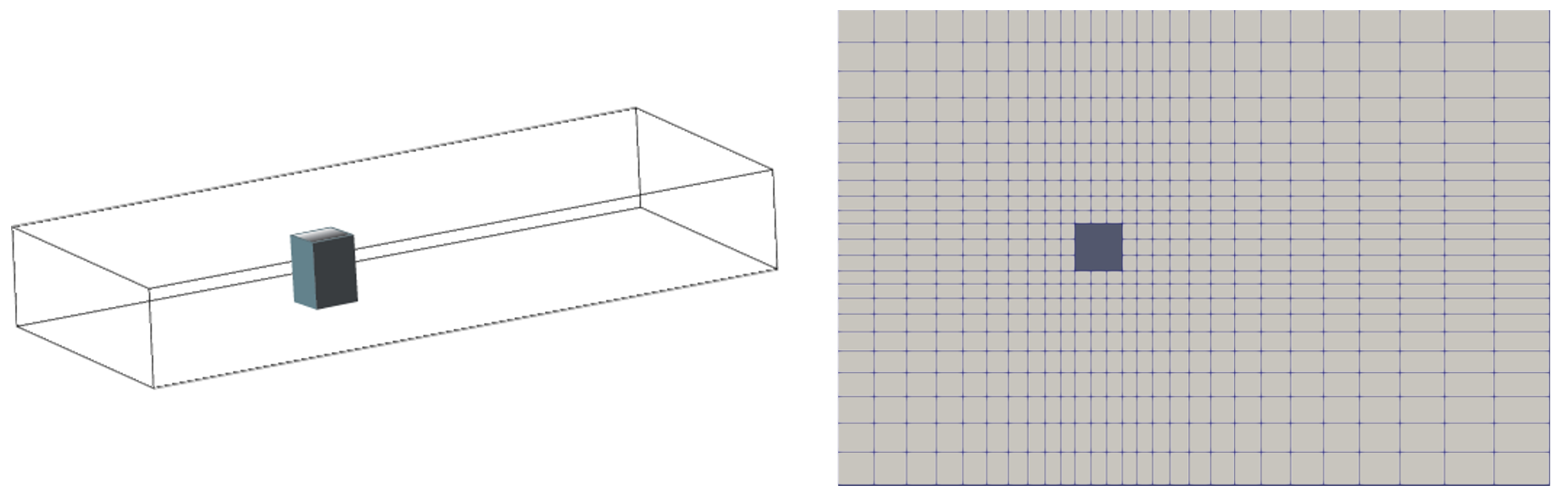
Fig. 4 Test model and its mesh
Setup agave
This script shows how to import the agave client, get the authorization (assuming that the user is already logged into DesignSafe), and access the OpenFOAM. Also
from agavepy.agave import Agave
ag = Agave.restore()
### see user profile
ag.profiles.get()
### access OpenFOAM
app = ag.apps.get(appId = 'openfoam-7.0u4')Submit the OpenFOAM job to DesignSafe/TACC
To utilize parallel computing for faster computation, the simulation is run in the DesignSafe workspace using the Texas Advanced Computing Center (TACC) computing resources. The following script shows how to set up the OpenFOAM configuration to run, and its submission to TACC, and check the status of the submitted job [3].
### Creating job file
jobdetails = {
# name of the job
"name": "OpenFOAM-Demo",
# OpenFOAM v7 is used in this use case
"appId": "openfoam-7.0u4",
# total run time on the cluster (max 48 hrs)
"maxRunTime": "00:02:00",
# the number of nodes and processors for parallel computing
"nodeCount": 1,
"processorsPerNode": 2,
# simulation results will be available in the user's archive directory
"archive": True,
# default storage system
"archiveSystem": "designsafe.storage.default",
# parameters for the OpenFOAM simulation
"parameters": {
# running blockmesh and/or snappyHexMesh (On)
"mesh": "On",
# running in parallel (On) or serial (Off)
"decomp": "On",
# name of OpenFOAM solver: pisoFoam is used in this use case
"solver": "pisoFoam"
},
"inputs": {
### directory where OpenFOAM files are stored: the path for the DH1_run directory in this use case
### check the web browser's URL at the DH1_run and use the path after "agave/" which includes one's USERNAME
### If DH1_run is located at Data depot > My Data > JupyterCFD > DH1_run, then the URL looks like:
### https://www.designsafe-ci.org/data/browser/agave/designsafe.storage.default/USERNAME/JupyterCFD/DH1_run
"inputDirectory": "agave://designsafe.storage.default/USERNAME/JupyterCFD/DH1_run"
}
}
### Submit the job to TACC
job = ag.jobs.submit(body=jobdetails)
### Check the job status
from agavepy.async import AgaveAsyncResponse
asrp = AgaveAsyncResponse(ag,job)
asrp.status
### if successfully submitted, then asrp.status outputs 'ACCEPTED'Post-processing on DesignSafe
Simulation results are stored in the Data Depot in the DesignSafe and available to be post-processed by users: Data Depot > My Data > archive > jobs. To visualize the flow fields, users can utilize a visualization tool in the Tools & Application menu in the DesignSafe (Fig. 5), Paraview [4], which can read OpenFOAM files using .foam file. Fig. 6 shows an example of the post-processing of simulation results in the Paraview.

Fig. 5 Paraview software in the DesignSafe
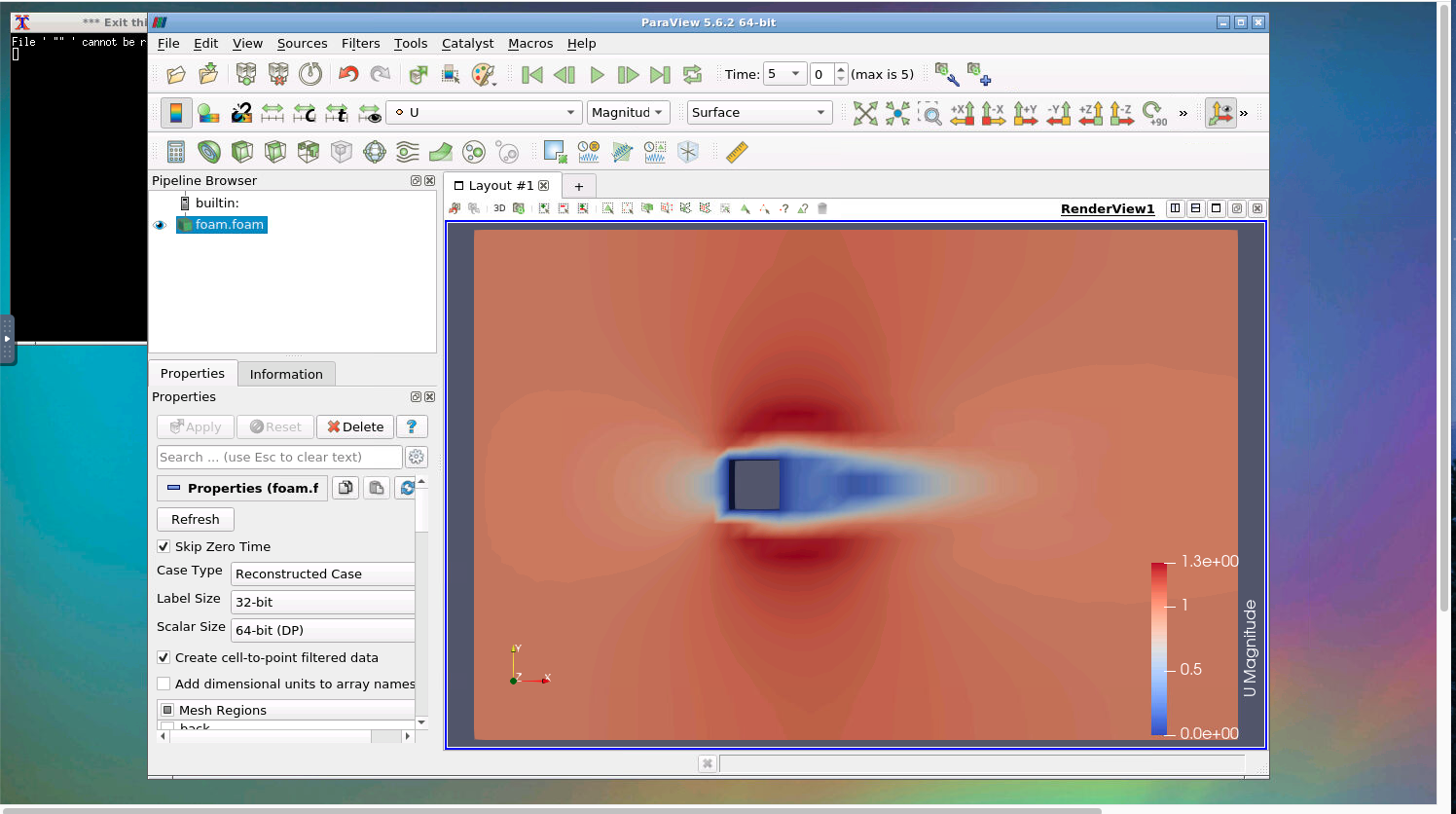
Fig. 6 Visualization of simulation results using the Paraview
For data analysis such as plotting the time series of drag or lift force coefficients, users can make a script in a Jupyter Notebook to load simulation results and make output figures using a python graphic library such as Matplotlib, etc. An example script using Matplotlib can also be found in this use case Jupyter Notebook.
References
[1] H. Jasak, A. Jemcov, Z. Tukovic, et al. OpenFOAM: A C++ library for complex physics simulations. In International workshop on coupled methods in numerical dynamics, volume 1000, pages 1-20. IUC Dubrovnik Croatia, 2007.
[2] OpenFOAM wiki. Pyfoam. https://openfoamwiki.net/index.php/Contrib/PyFoam. Online; accessed 24-Feb-2022.
[3] Harish, Ajay Bangalore; Govindjee, Sanjay; McKenna, Frank. CFD Notebook (Beginner). DesignSafe-CI, 2020.
[3] N. Vuaille. Controlling paraview from jupyter notebook. https://www.kitware.com/paraview-jupyter-notebook/. Online; accessed 24-Feb-2022.
Seismic Use Cases
- Site Response Analysis and Model Calibration (OpenSees, SimCenter quoFEM, Jupyter, HPC)
- Simulating the Seismic Performance of Reinforced Concrete Walls (OpenSees, Jupyter, HPC)
- Soil-Structure-Interaction Simulations (OpenSees, STKO, Jupyter, HPC)
- Experimental Visualization of Shaking Table Data (Jupyter, Interactive Data Analysis, UCSD NHERI Facility)
Field Sensing Wind Events
Field Sensing Wind Events
Wind Data Analysis Tools
Soundarya Sridhar - Florida Institute of Technology
Jean-Paul Pinelli - Florida Institute of Technology
M.A. Ajaz - Florida Institute of Technology
Key Words: wireless sensors network, Tapis, Plotly, and Hurricane Ian
Florida Tech (FIT) teams deploy networks of wireless sensors on residential houses during high impact wind events or on full scale wind tunnel models. Each deployment might include pressure, temperature and humidity sensors alongside different anemometers and a conical scanning infrared LIDAR. Figure 1 describes the workflow, which starts with uploading the data to DesignSafe through authentication tokens created in Tapis. Once on DesignSafe, three Jupyter notebooks process and visualize the instruments data for analyses. The notebooks provide a user friendly and interactive environment that can adapt to different datasets. For this project, the notebooks perform quasi static real-time analysis, assess sensor performance, and study pressure variations for different wind conditions and data correlation. The user interactivity of these notebooks facilitates an easy adaptation to different datasets with little to no-change in code.
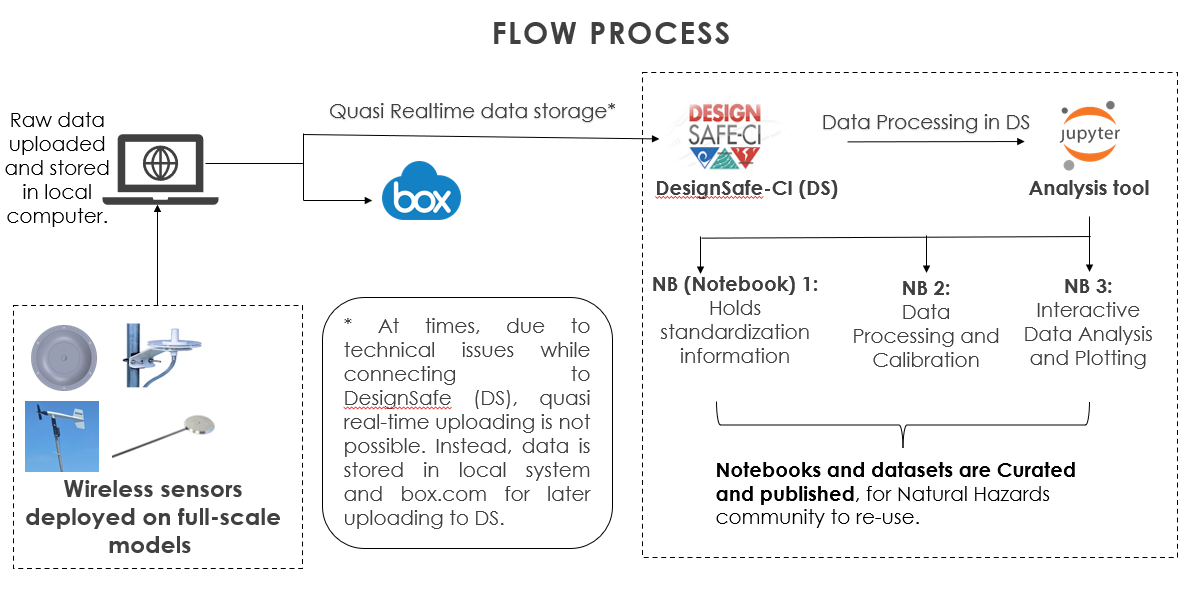
Figure 1. Workflow
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
Jupyter Notebooks for WOW Sliding Patio Doors
Jupyter Notebooks for WOW Storm Shield
DesignSafe Resources
The following DesignSafe resources were used in developing this use case.
- Jupyter notebooks on DS Juypterhub
- Subramanian, C., J. Pinelli, S. Lazarus, J. Zhang, S. Sridhar, H. Besing, A. Lebbar, (2023) "Wireless Sensor Network System Deployment During Hurricane Ian, Satellite Beach, FL, September 2022", in Hurricane IAN Data from Wireless Pressure Sensor Network and LiDAR. DesignSafe-CI. https://doi.org/10.17603/ds2-mshp-5q65
- Video Tutorial (Timestamps - 28:01 to 35:04): https://youtu.be/C2McrpQ8XmI?t=1678
Background
Citation and Licensing
-
Please cite Subramanian et al. (2022), Pinelli et al. (2022), J. Wang et al. (2021) and S. Sridhar et al. (2021) to acknowledge the use of any resources from this use case.
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License.
Description
Implementaton
Quasi-real time Data Upload with Tapis
The user needs a DesignSafe-CI (DS) account. During deployment, data is uploaded to DS in user defined time interval. Tapis CLI and Python 3 executable enable this feature and must be installed on the local system. The user initiates Tapis before every deployment through Windows PowerShell and Tapis creates a token as described below:
Video Tutorial (Timestamps - 28:01 to 35:04): https://www.youtube.com/watch?v=C2McrpQ8XmI
User Guide
- Turn on Windows Power Shell and enter the command tapis auth init -interactive.
- Enter designsafe for the tenant name.
- Enter the DesignSafe username and password of the authorized user.
- Choose to set up Container registry access and Git server access or skip this step by pressing the return key.
- Create a token using the command tapis auth tokens create. At the end, the response will appear on the cmd line as shown in Figure 2
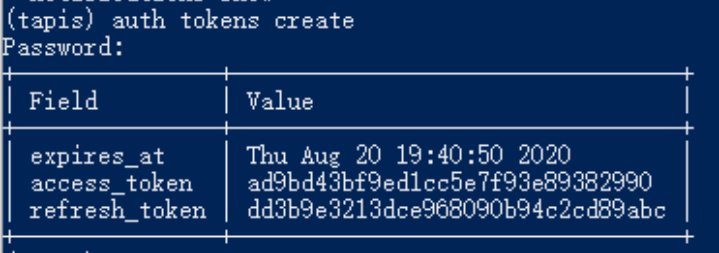
Figure 2.
Using Jupyter Notebooks
Instructions
Using JupyterHub on DesignSafe
Accessing JupyterHub
Navigate to the JupyterHub: Use this link to go directly to the JupyterHub portal on DesignSafe. Sign In: You must have a TACC (Texas Advanced Computing Center) account to access the resources. If you do not have an account, you can register here. Access the Notebook: Once signed in, you can access and interact with the Jupyter notebooks available on your account. To run this Project, you must copy it to your MyData directory to make it write-able as it is read only in NHERI- published directory. Use your favorite way to lunch a Jupyter Notebook and then open the FirstMap.ipynb file.
-
Run the following command cell to copy the project to your MyData or change path to wherever you want to copy it to: after opening this Notebook in MyData you don't have to run the below cell again !umask 0022; cp -r/home/jupyter/NHERI-Published/PRJ-4535v2 /home/jupyter/MyData/PRJ-4535; chmod -R u+rw /home/jupyter/MyData/PRJ-4535
-
Navigate to your 'MyData' directory. For illustrative purposes, input files have been created and shared in this project. These files have been pre-processed and conveniently organized used to illustrate the data collection, integration, and visualization on the map. The outcomes as follows:
- CB_WSNS_WOW_6-22-21: This folder contains
a. Calibration Constants_WSNS_WOW_6-22-21_ALL.csv file.
b. Standardization_Info_FITWSNS_WOW_6-22-21.csv file. c. CSV files and pkl files. - html_images: input and output are saved as html_images used are included in this folder
- Res.csv : contains, Sensor, WS (MPH),WD (deg), Min, Max, Mean (mbar), Stddev
- RW_WOW_6-21-2021_SlidingPatioDoors_WSNS
- Jupyter Notebooks for WOW_Sliding Patio Doors
a. WOW_6-22-21_NB1__Standardization File.ipynb
b. WoW_6-22-21_NB2_WSNS POST PROCESSING.ipynb c. WoW_6-22-21_NB3_INTERACTIVE ANALYSIS.ipynb d. Box.jpg, SensorLoc_Glass Slider_6_22.jpg, Sliders.jpg.
- CB_WSNS_WOW_6-22-21: This folder contains
a. Calibration Constants_WSNS_WOW_6-22-21_ALL.csv file.
To save time and memory, the project uses three different notebooks. For any event, either a field deployment or a wind tunnel experiment, the first notebook inputs metadata (sensor information, data columns, timestamp formats) for the dataset and is ideally used once for every event. It outputs a csv file containing the metadata required to run the second notebook. The second notebook calibrates raw data and organizes them into csv and pickled files. This notebook may be run more than once depending on how often new data is uploaded during the event. With the third notebook, users analyse and visualize the data interactively. This is the most frequently used notebook and is run every time the data needs to be analysed. There is no need to execute the notebooks sequentially every time an analysis is done. Figure 3 below illustrates the possible sequences of analysis:
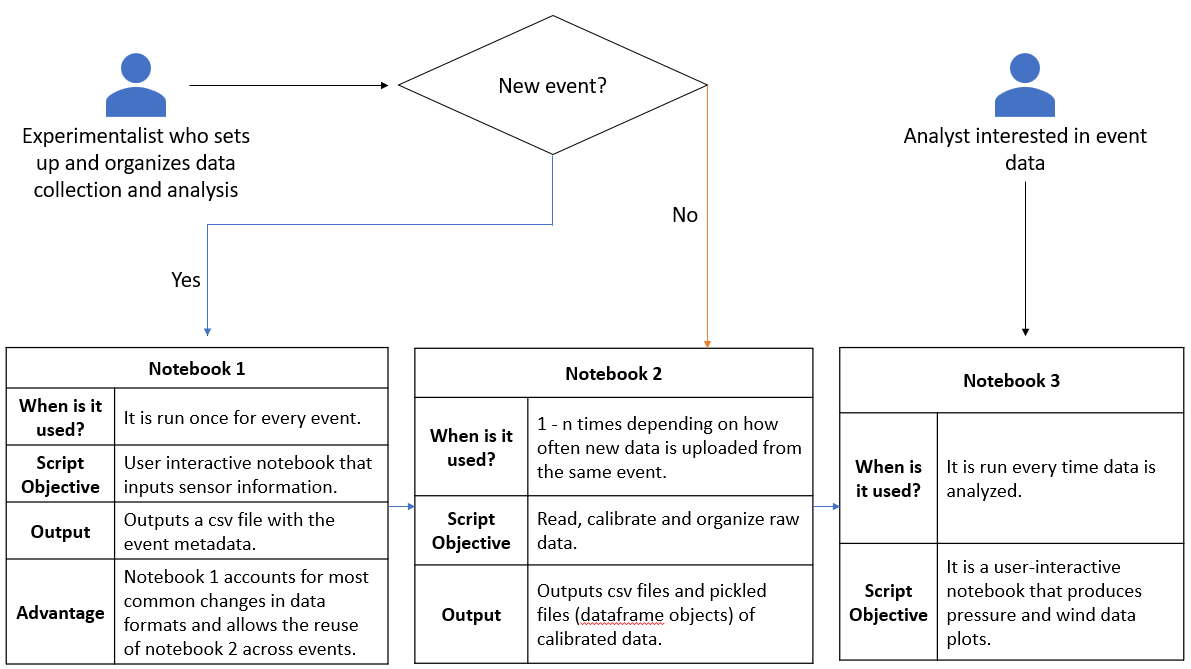
Figure 3. Sequence of analysis
Adaptation to Different Datasets
The first notebook is a user interactive guide to input important raw data information. This notebook saves time as the user does not have to read, understand and edit the code to change information regarding sensors, columns and data formats. For example, WSNS deployment during the tropical storm Isaias (8/2/2020) used an old and a new WSNS system. The first notebook documented the significant differences in data storage between the two systems. This accelerates data processing as there is no change required in code and the file generated by the notebook acts as a metadata for the second notebook responsible for data processing. Figure 4 below shows snapshots of the output file created by the first notebook describing raw data information from two different systems.
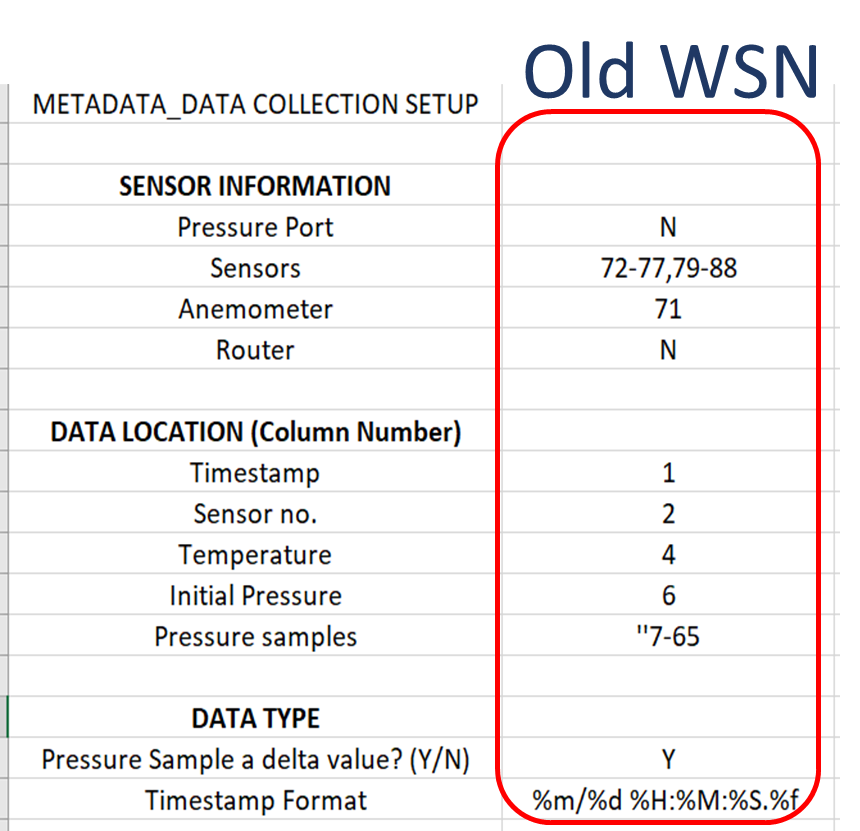
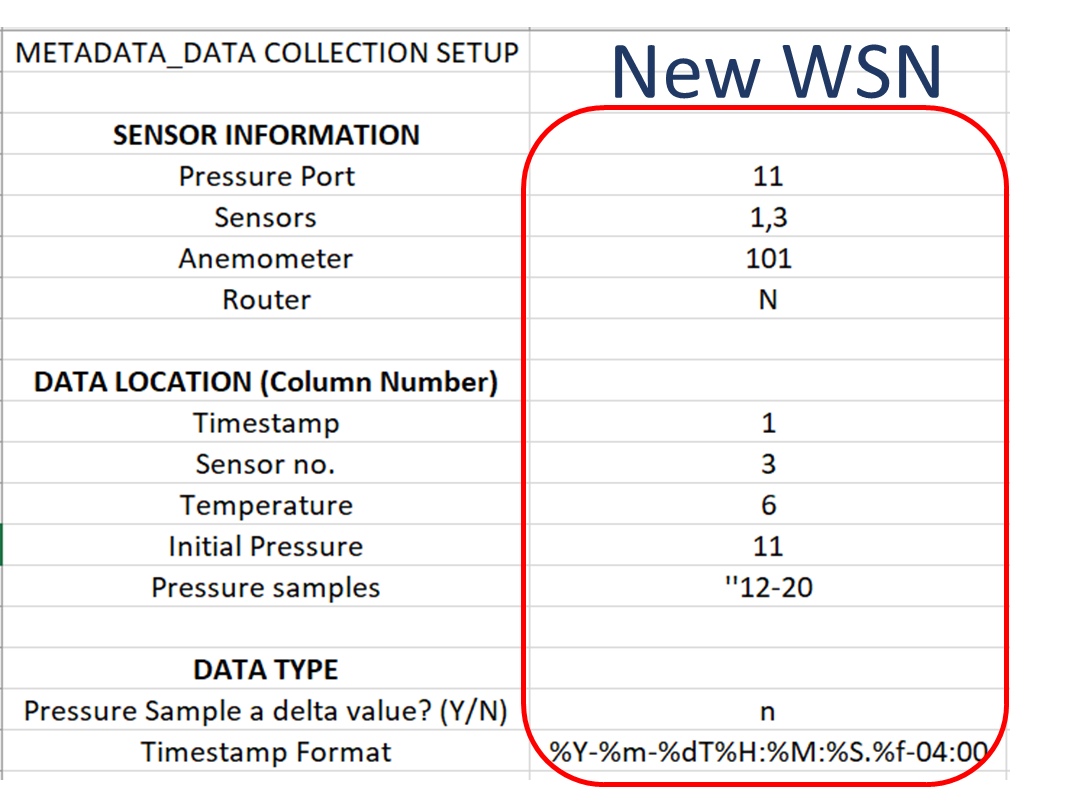
Figure 4. Snapshots of output files
Jupyter Notebooks for WOW Sliding Platio Door
Analyses Notebooks and Examples
The project goal is to measure pressure variation on non-structural components during either strong wind events or full-scale testing in the WoW, using the network of wireless sensors. The analysis notebooks on DesignSafe are user interactive with markdowns describing the test. They also provide the users with several options to visualize the data. For example, figure 5a shows the analysis notebook for Isaias (tropical storm on August 1-3, 2020) while Figure 5b shows the analysis notebook for WoW test. The markdowns have important information and pictures from the deployment, and instructions for the user to easily access data.
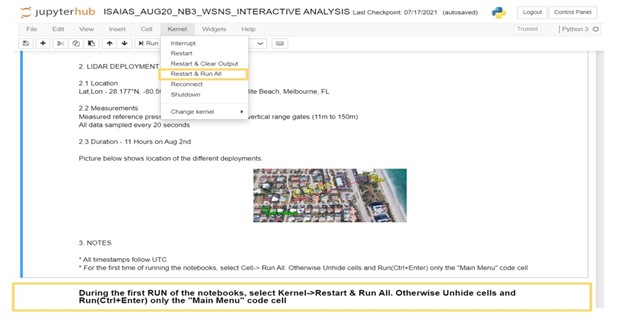
Figure 5a. Analysis notebook for Isaias (field deployment)
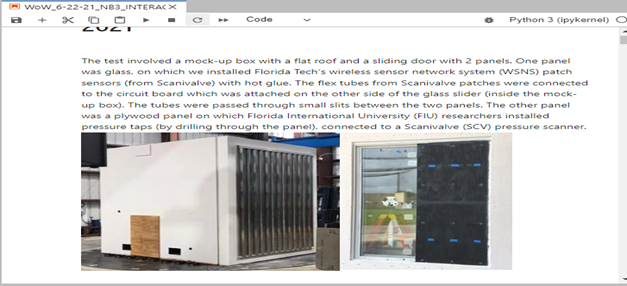
Figure 5b. Analysis notebook for WoW test (wind tunnel deployment)
Figure 6 shows the menu that allows users to select from options and look at specific time windows or test conditions.
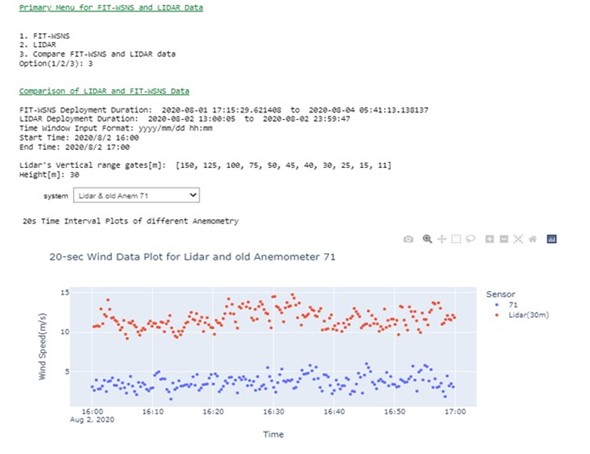
Figure 6. Menu options in Analysis Notebook
Using Plotly for Data Driven Animation Frames
The project objective is to study high impact wind events on non-structural components of residential houses. After the deployment or the test, Jupyter notebooks process and visualize important data for different purposes, including among others: comparisons to ASCE 7 standard; and assessment of sensor performance with respect to wind conditions. Plotly can create animation frames to look at a snapshot of data from all sensors in different test conditions or even at different timestamps. A single line of code enabled with the right data frame can quickly reveal trends in the data and facilitate troubleshooting of any system errors. Figure 7 below shows an application of plotly for one of the Wall of Wind tests for glass sliding doors. The test model was a mock-up box with flat roof, and full-scale glass sliding doors, which were tested at 105 mph for different wind directions. At uniform velocity, data for each wind direction was collected for 3 minutes and the program computed pressure coefficient Cp values averaged over that time window. A 2D scanner plot was created with x and z dimensions with each point representing a sensor whose colour corresponded to a Cp value on the colour scale. A single line of code enables the animation frame, which reveals important information:
px.scatter(dataframe, x=x column, y=y column, color=scatter point values, text=text to be displayed for each point, range_color=color scale range, animation_frame=variable for each animation frame, title = plot title)Including dimensions and trace lines to the plots can add more clarity.

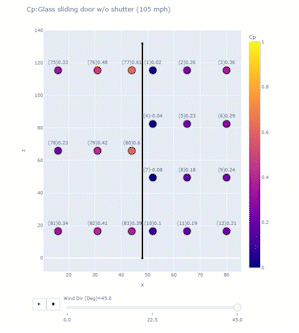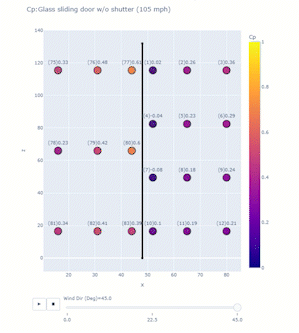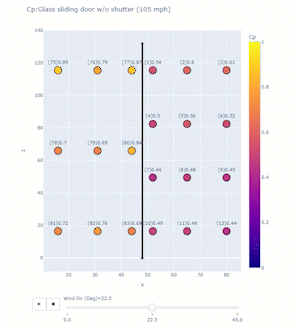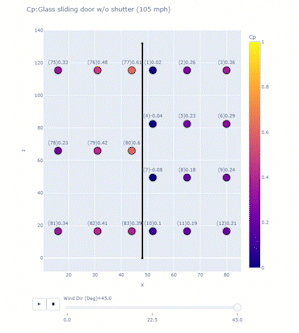
Figure 7. Application of Plotly for one of the Wall of Wind tests for glass sliding doors
Plotly features example
The exercise below is an illustration of these plotly features:
Requirements:
Access Jupyter Notebook on DesignSafe. Once you have your notebook open and you don’t have plotly dash installed, go ahead and use: !pip install dash==1.14.0 --user
Building the Dataframe: Building the Dataframe: Consider a box of spheres that change their numbers ranging from 1 to 10 every hour. You want to look at how the number changes for 12 hours.
Code
#import Libraries
import random
import pandas as pd
# Define necessary columns
spheres = [1, 2, 3, 4, 5]
x = [6, 14, 10, 6, 14]
y = [6, 6, 10, 14, 14]
rad = []
# Generating 5 random numbers ranging from 1 to 10 for the first hour
for i in range(0, 5):
n = random.randint(1, 10)
rad.append(n)
hour = 1
Label = ['1', '2', '3', '4', '5']
# DataFrame for the first hour
df = pd.DataFrame(spheres, columns=['Sphere'])
df['x'] = x
df['y'] = y
df['number'] = rad
df['hour'] = hour
df['label'] = Label
# Loop for the next 11 hours
for i in range(0, 11):
hour = hour+1
temp = pd.DataFrame(spheres, columns=['Sphere'])
temp['x'] = x
temp['y'] = y
rad = []
for i in range(0, 5):
n = random.randint(1, 10)
rad.append(n)
temp['number'] = rad
temp['hour'] = hour
temp['label'] = Label
df = df.append(temp, ignore_index=True)
print(df)Matching the right columns to suit the syntax will result in an animation frame and a slider!
import plotly.express as px
import plotly.graph_objects as go
from IPython.display import display, HTML
fig = px.scatter(df, x='x',y='y', color='number',text="label",
animation_frame='hour',title='Magic Box') #animation frame
fig.update_traces(textposition='top center',mode='markers', marker_line_width=2, marker_size=40)
trace1 = go.Scatter(x=[2, 2], y=[2, 18],line=dict(color='black', width=4),showlegend=False) #Tracelines to create the box
trace2 = go.Scatter(x=[2, 18], y=[18, 18],line=dict(color='black', width=4),showlegend=False)
trace3 = go.Scatter(x=[18, 18], y=[18, 2],line=dict(color='black', width=4),showlegend=False)
trace4 = go.Scatter(x=[18, 2], y=[2, 2],line=dict(color='black', width=4),showlegend=False)
fig.add_trace(trace1)
fig.add_trace(trace2)
fig.add_trace(trace3)
fig.add_trace(trace4)
fig.update_layout(autosize=False,width=500,height=500,showlegend=True)
html_file_path11 = parent_dir+'/html_images/magic_box.html'
fig.write_html(html_file_path11, include_plotlyjs='cdn')
display(HTML(filename=html_file_path11))display(HTML(filename=html_file_path11))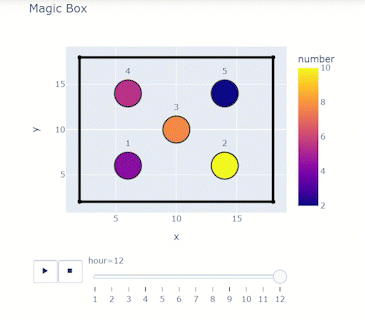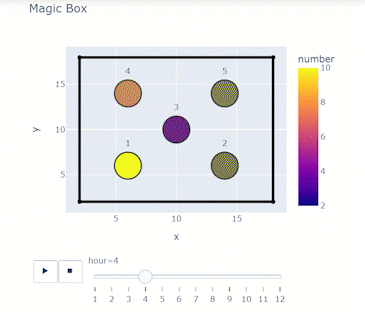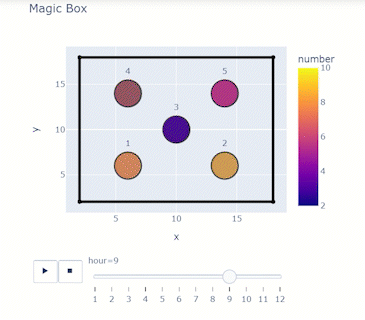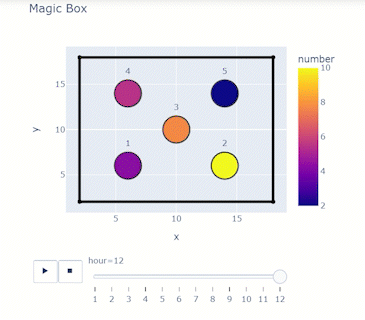

OpenSees Model Calibration
OpenSees Model Calibration
From Constitutive Parameter Calibration to Site Response Analysis
Pedro Arduino - University of Washington
Sang-Ri Yi - SimCenter, UC Berkeley
Aakash Bangalore Satish - SimCenter, UC Berkeley
Key Words: quoFEM, OpenSees, Tapis, Python
A collection of educational notebooks to introduce model-parameter calibration and site response analysis using OpenSees in DesignSafe-CI.
Resources
Jupyter Notebooks
The following Jupyter notebooks are made available to facilitate the analysis of each case. They are described in detail in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
| Site Response | Notebook |
|---|---|
| FreeField Response | freeFieldJupyterPM4Sand_Community.ipynb |
| quoFEM | Notebook |
|---|---|
| Sensitivity analysis | quoFEM-Sensitivity.ipynb |
| Bayessian calibration | quoFEM-Bayesian.ipynb |
| Forward propagation | quoFEM-Propagation.ipynb |
DesignSafe Resources
The following DesignSafe resources were used in developing this use case.
- DesignSafe - Jupyter notebooks on DS Juypterhub
- SimCenter - quoFEM
- Simulation on DesignSafe - OpenSees
Background
Citation and Licensing
-
Please cite Aakash B. Satish et al. (2022) to acknowledge the use of resources from this use case.
-
Please cite Sang-Ri Yi et al. (2022) to acknowledge the use of resources from this use case.
-
Please cite Chen, L. et al. (2021) to acknowledge the use of resources from this use case.
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License.
Description
Seismic site response refers to the way the ground responds to seismic waves during an earthquake. This response can vary based on the soil and rock properties of the site, as well as the characteristics of the earthquake itself.
Site response analysis for liquefiable soils is fundamental in the estimation of demands on civil infrastructure including buildings and lifelines. For this purpose, current state of the art in numerical methods in geotechnical engineering require the use of advance constitutive models and fully couple nonlinear finite element (FEM) tools. Advanced constitutive models require calibration of material parameters based on experimental tests. These parameters include uncertainties that in turn propagate to uncertenties in the estimation of demands. The products included in this use-case provide simple examples showing how to achieve site response analysis including parameter identification and uncertainty quantification using SimCenter tools and the DesignSafe cyber infrastructure.
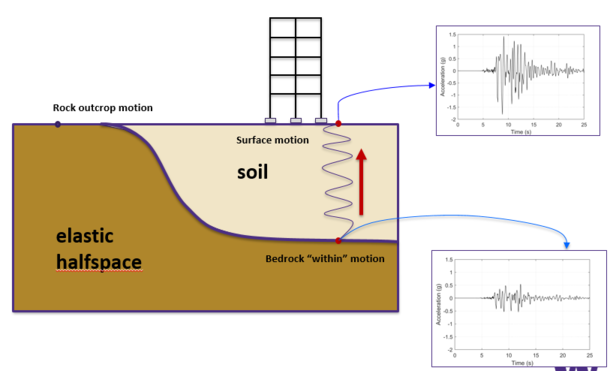
Fig.1 - Site response problem
Implementation
This use-case introduces a suite of Jupyter Notebooks published in DesignSafe that navigate the process of constitutive model parameter calibration and site response analysis for a simple liquefaction case. They also introduce methods useful when using DesignSafe infrastructure in TACC. All notebooks leverage existing SimCenter backend functionality (e.g. Dakota, OpenSees, etc) implemented in quoFEM and run locally and in TACC through DesignSafe. The following two pages address these aspects, including:
-
Site response workflow notebook: This notebook introduces typical steps used a seismic site response analysis workflow taking advantage of Jupyter, OpenSees, and DesignSafe-CI.
-
quoFEM Notebooks: These notebooks provide an introduction to uncertainty quantification (UQ) methods using quoFEM to address sensitivity, Bayesian calibration, and forward propagation specifically in the context of seismic site response. Three different analysis are discussed including:
a. Global sensitivity analysis This notebook provides insight into which model parameters are critical for estimating triggering of liquefaction.
b. Parameter calibration notebook: This notebook is customized for the PM4Sand model and presents the estimation of its main parameters that best fit experimental data as well as their uncertainty.
c. Propagation of parameter undertainty in site response analysis notebook: This notebook introduces methods to propagate material parameter uncertainties in site reponse analysis.
Seismic Response of Concrete Walls
Seismic Response of Concrete Walls
Modeling Reinforced Concrete Walls using Shell Elements in OpenSees and Using Jupyter to Post Process Results
Josh Stokley - University of Washington
Laura Lowes - University of Washington
Key Words: OpenSees, Jupyter, HPC
The purpose of this use case is to be able to model, simulate, and post process multiple reinforced concrete walls at once. This use case uses jupyter notebooks to model these walls with shell elements and uses OpenSeesMP on DesignSafe to simulate the models. The documentation of this use case will use a single wall, RW1, as an example to understand the workflow and objectives of this use case.
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
DesignSafe Resources
The following DesignSafe resources were used in developing this use case.
Background
Citation and Licensing
-
Please cite Shegay et al. (2021) to acknowledge the use of any data from this use case.
-
Please cite Lu XZ et al. (2015) to acknowledge the use of the modeling strategy from this use case.
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License .
Description
Data
The walls that are modeled are defined in a database provided by Alex Shegay. The database is a MATLAB variable of type 'structure'. The tree-like structure of the variable consists of several levels. Each level consists of several variables, each being a 1x142 dimension array. Each entry within the array corresponds to a separate wall specimen. The order of these entries is consistent throughout the database and reflects the order of walls as appearing in the 'UniqueID' array.
Modeling
The modeling techniques are inspired by the work of Lu XZ. The modeling of these walls make use of the MITC4 shell element. This element smears concrete and steel in multiple layers through the thickness of the element. Figure 1 demonstrates this. Within the shell element, only the transverse steel is smeared with the concrete. The shell elements are modeled to be square or close to square for best accuracy, an assumption that follows this is that cover concrete on the ends of the wall are not taken into account as it would produce skinny elements that would cause the wall to fail prematurely. The vertical steel bars are modeled as trusses up the wall to better simulate the stress of those bars. The opensees material models that are used are:
-
PlaneStressUserMaterial- Utilizes damage mechanisms and smeared crack model to define a multi-dimensional concrete model
- Variables include: compressive strength, tensile strength, crushing strength, strain at maximum and crushing strengths, ultimate tensile strain, and shear retention factor
- Model can be found in Lu XZs citation
-
Steel02- Uniaxial steel material model with isotropic strain hardening
- Variables include: yield strength, initial elastic tangent, and strain hardening ratio
- Model can be found here: Steel02 OpenSees
Figure 1: Smeared shell element representation
Example
RW1 is modeled from the database to produce a tcl file that represents the geometry, material, and simulation history of the wall. The wall is 150 inches high, 46.37 inches long, and 4 inches thick. It consists of 1292 amount of nodes, 1200 amount of shell elements, and 900 amount of steel truss elements. MITC4 shell elements are used to smear the concrete and transverse steel into the thickness while the vertical reinforce bars are modeled as truss elements. RW1 had a compression buckling failure mode in the lab. More information on RW1 and its experimental results can be found here: Wallace et al. (2004)
The use case workflow involves the following steps:
- Using Jupyter notebook modeling script to create input file for OpenSees
- Running input file through HPC on DesignSafe
- Using Jupyter notebook post processing scripts to evaluate model
Create Input File using Modeling Script
The modeling script is broken up into 2 notebooks, the first notebook imports the variables to build the wall into an array. The second notebook builds out the tcl file that will be ran through openseees. The sections defined below are from the second notebook.
The matlab to python script can be found here: Matlab_to_Python.ipynb
The jupyter notebook that creates the OpenSees input file can be found here: TCL_Script_Creator.ipynb
Reinforced Concrete Wall Database
Each wall in the database has a number corresponding to its unique ID. This number will be the single input to the modeling script to create the script. The use case will loop through multiple numbers to create multiple files at once and run them through opensees. Variables are separated in the database by sections. For example, under the section 'Geometry', one can find the heights of the walls, the thickness of walls, the aspect ratios, and so on. By parsing through these sections, the necessary information can found and imported into the modeling script to build out the wall.
RW1 is wall 33 in the database (with the first wall index starting at 0) and using that index number, the modeling script can grab everything that defines RW1.
Modeling Script
The sections of the modeling script are:
Section 1: Initialization of the model
- The degrees of freedom and the variables that carry uncertainty are defined.
Section 2: Defines nodal locations and elements
- Nodes are placed at the locations of the vertical bars along the length of the wall.
- If the ratio of the length of the wall to the number of elements is too coarse of a mesh, additional nodes are placed in between the bars.
- The height of each element is equal to the length of the nodes in the boundary to create square elements up the wall.
Section 3: Defines material models and their variables
- The crushing energy and fracture energy are calculated and wrote to the .tcl file. The equations for these values come from (Nasser et al. (2019) ) Below is the code:
self.gtcc = abs((0.174*(.5)**2-0.0727*.5+0.149)*((self.Walldata[40]*1000*conMult)/1450)**0.7) #tensile energy of confined
self.gtuc = abs((0.174*(.5)**2-0.0727*.5+0.149)*((self.Walldata[40]*1000)/1450)**0.7) # tensile energy of unconfined
self.gfuc = 2*self.Walldata[40]*6.89476*5.71015 #crushing energy of unconfined
self.gfcc = 2.2*self.gfuc #crushing energy of confined- The crushing strain (epscu) and fracture strain (epstu) can then be calculated from the energy values.
- The material models are then defined.
- The concrete material opensees model: nDmaterial PlaneStressUserMaterial $matTag 40 7 $fc $ft $fcu $epsc0 $epscu $epstu $stc.
- 'fc' is the compressive strength, 'ft' is the tensile strength, 'fcu' is the crushing strength, and 'epsc0' is the strain at the compressive strength.
- The steel material opensees model: uniaxialMaterial Steel02 $matTag $Fy $E $b $R0 $cR1 $cR2.
- 'Fy' is the yield strength, 'E' is the youngs modulus, 'b' is the strain hardening ratio, and 'R0', 'cR1', and 'cR2' are parameters to control transitions from elastic to plastic branches.
- minMax wrappers are applied to the steel so that if the steel strain compresses more than the crushing strain of the concrete or exceeds the ultimate strain of the steel multiplied by the steel rupture ratio, the stress will go to 0.
Section 4: Defines the continuum shell model
- The shell element is split up into multiple layers of the cover concrete, transverse steel, and core concrete.
- The cover concrete thickness is defined in the database, the transverse steel thickness is calculated as:
- total layers of transverse steel multiplied by the area of the steel divided by the height of the wall.
- The total thickness of the wall is defined in the database so after the cover concrete and steel thicknesses are subtracted, the core concrete takes up the rest.
Section 5: Defines the elements
- The shell element opensees model is: element ShellMITC4 $eleTag $iNode $jNode $kNode $lNode $secTag
- 'eleTag' is the element number, the next four variables are the nodes associated to the element in ccw, and 'secTag' is the section number that defines the thickness of the element.
- There are usually two sections that are defined, the boundary and the web. Based on how many nodes are in the boundary, the script will print out the elements for the left side of the boundary, then for the entire web region, and lastly for the right side of the boundary. This process is repeated until the elements reach the last row of nodes.
- For the vertical steel bars, the truss element opensees model is used: element truss $eleTag $iNode $jNode $A $matTag.
- The node variables are defined as going up the wall so if a wall has 10 nodes across the base, the first truss element would connect node 1 to node 11.
- 'A' is the area of the bar and 'matTag' is the material number applied to the truss element.
- The script prints out truss elements one row at a time so starting with the left furthest bar connecting to each node until the height of the wall is reached and then next row is started the bar to the right.
Section 6: Defines constraints
- The bottom row of nodes are fixed in all degrees of freedom.
Section 7: Defines recorders
- The first two recorders capture the force reactions in the x-direction of the bottom row of nodes and the displacements in the x-direction of the top row of nodes. These recorders will be used to develop load-displacement graphs.
- The next eight recorders capture stress and strain of the four gauss points in the middle concrete fiber of all the elements and store them in an xml file. These recorders will be used to develop stress and strain profile movies, give insight to how the wall is failing, and how the cross section is reacting.
- The last two recorders capture the stress and strain of all the truss elements. These will be used to determine when the steel fails and when the yield strength is reached.
- Recorders are defined as below where 'firstRow' and 'last' are the nodes along the bottom and top of the wall 'maxEle' is the total amount of shell elements and 'trussele' is the total elements of shell elements and truss elements in the wall.
self.f.write('recorder Node -file baseReactxcyc.txt -node {} -dof 1 reaction\n'.format(' '.join(firstRow)) )
self.f.write('recorder Node -file topDispxcyc.txt -node {} -dof 1 disp \n'.format(' '.join(last)))
self.f.write('recorder Element -xml "elementsmat1fib5sig.xml" -eleRange 1 ' + str(self.maxEle) + ' material 1 fiber 5 stresses\n')
self.f.write('recorder Element -xml "elementsmat2fib5sig.xml" -eleRange 1 ' + str(self.maxEle) + ' material 2 fiber 5 stresses\n')
self.f.write('#recorder Element -xml "elementsmat3fib5sig.xml" -eleRange 1 ' + str(self.maxEle) + ' material 3 fiber 5 stresses\n')
self.f.write('#recorder Element -xml "elementsmat4fib5sig.xml" -eleRange 1 ' + str(self.maxEle) + ' material 4 fiber 5 stresses\n')
self.f.write('recorder Element -xml "elementsmat1fib5eps.xml" -eleRange 1 ' + str(self.maxEle) + ' material 1 fiber 5 strains\n')
self.f.write('recorder Element -xml "elementsmat2fib5eps.xml" -eleRange 1 ' + str(self.maxEle) + ' material 2 fiber 5 strains\n')
self.f.write('#recorder Element -xml "elementsmat3fib5eps.xml" -eleRange 1 ' + str(self.maxEle) + ' material 3 fiber 5 strains\n')
self.f.write('#recorder Element -xml "elementsmat4fib5eps.xml" -eleRange 1 ' + str(self.maxEle) + ' material 4 fiber 5 strains\n')
self.f.write('recorder Element -xml "trusssig.xml" -eleRange ' + str(self.maxEle+1) + ' ' + str(self.trussele)+ ' material stress\n')
self.f.write('recorder Element -xml "trussseps.xml" -eleRange ' + str(self.maxEle+1) + ' ' + str(self.trussele)+ ' material strain\n')Section 8: Defines and applies the gravity load of the wall
- The axial load of the wall is defined in the database and distributed equally amongst the top nodes and a static analysis is conducted to apply a gravity load to the wall.
Section 9: Defines the cyclic analysis of the wall
- The experimental displacement recording of the wall is defined in the database and the peak displacement of each cycle is extracted.
- If the effective height of the wall is larger than the measured height, a moment is calculated from that difference and uniformly applied in the direction of the analysis to each of the top nodes.
- The displacement peaks are then defined in a list and ran through an opensees algorithm.
- This algorithm takes each peak and displaces the top nodes of the wall by 0.01 inches until that peak is reached, it then displaces by 0.01 inches back to zero where it then takes on the next peak. This process continues until failure of the wall or until the last peak is reached.
The last section of this notebook creates a reference file that holds variables needed for postprocessing. In order, those variables are: * Total nodes along the width of the wall * Total nodes along the width of the wall that are connected to a truss element * Total nodes in the file * Total elements in the file * Displacement peaks in the positive direction * Fracture strength of the concrete * total layers of elements in the file * Unique ID of the wall * filepath to the folder of the wall * filepath to the tcl file
Running Opensees through HPC
(Script needs to be established on design safe. I have a working notebook, just need to connect it with modeling script)
Post Processing
After the script is finished running through OpenSees, there are multiple post-processing scripts that can be used to analyze the simulation and compare it to the experimental numbers.
Load-Displacement Graph
The Load-Displacement script compares the experimental cyclic load history to the simulated cyclic load output. The x axis is defined as drift % which is calculated as displacement (inches) divided by the height of the wall. The y axis is defined as shear ratio and calculated as force (kips) divided by cross sectional area and the square root of the concrete compressive strength. This Script can be found here: DisplacementLoadHistory.ipynb
Cross Sectional Analysis of Concrete and Steel
The cross sectional script shows stress and strain output across the cross section of the first level for the concrete and steel at various points corresponding with the positive displacement peaks. This script can be found here: CrossSectionSteelConcreteProfile.ipynb


Stress and Strain Profile Movies
The Stress/Strain profile movie script utilizes plotly to create an interactive animation of stresses and strains on the wall throughout the load history. The stress animations are vertical stress, shear stress, and maximum and minimum principal stress. The strain animations are vertical strain, shear strain, and maximum and minimum principal strain. This script can be found here: Movies.ipynb

Crack Angle of Quadrature Points
The crack angle script will show at what angle each quadrature point cracks. When the concrete reaches its fracture strength in the direction of the maximum principal stress, it is assumed that it cracked and the orientation at that point is then calculated shown on the graph. The blue lines indicates the crack angle was below the local x axis of the element and the red line means the crack angle was above the local x axis of the element. This script can be found here: CrackedModel.ipynb

Soil Structure Interaction
Soil Structure Interaction
Integration of OpenSees-STKO-Jupyter to Simulate Seismic Response of Soil-Structure-Interaction
Yu-Wei Hwang - University of Texas at Austin
Ellen Rathje - University of Texas at Austin
Key Words: OpenSees, STKO, Jupyter, HPC
This use case example shows how to run an OpenSeesMP analysis on the high-performance computing (HPC) resources at DesignSafe (DS) using the STKO graphical user interface and a Jupyter notebook. The example also post-processes the output results using python scripts, which allows the entire analysis workflow to be executed within DesignSafe without any download of output.
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
| Scope | Notebook |
|---|---|
| Submit job to STKO-compatible OpenSees | SSI_MainDriver.ipynb |
| Post-Processing in Jupyter | Example post-processing scripts.ipynb |
DesignSafe Resources
The following DesignSafe resources were used in developing this use case.
Background
Citation and Licensing
-
Please cite Hwang et al. (2021) to acknowledge the use of any resources from this use case.
-
Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
-
This software is distributed under the GNU General Public License .
Description
A hypothetical three dimensional soil–foundation–structure system on liquefiable soil layer is analyzed using OpenSees MP. The soil profile first included a 12-m thick dense sand layer with Dr of 90%, followed by a 4-m thick loose sand layer with Dr of 40%, and overlaid by a 2-m thick dense sand layer. The ground water table was at ground surface. An earthquake excitation was applied at the bottom of the soil domain under rigid bedrock conditons. A three-story, elastic structure was considered on a 1-m-thick mat foundation. The foundation footprint size (i.e., width and length) was 9.6m x 9.6m with bearing pressure of 65 kPa. Additional information can be found in Hwang et al. (2021)
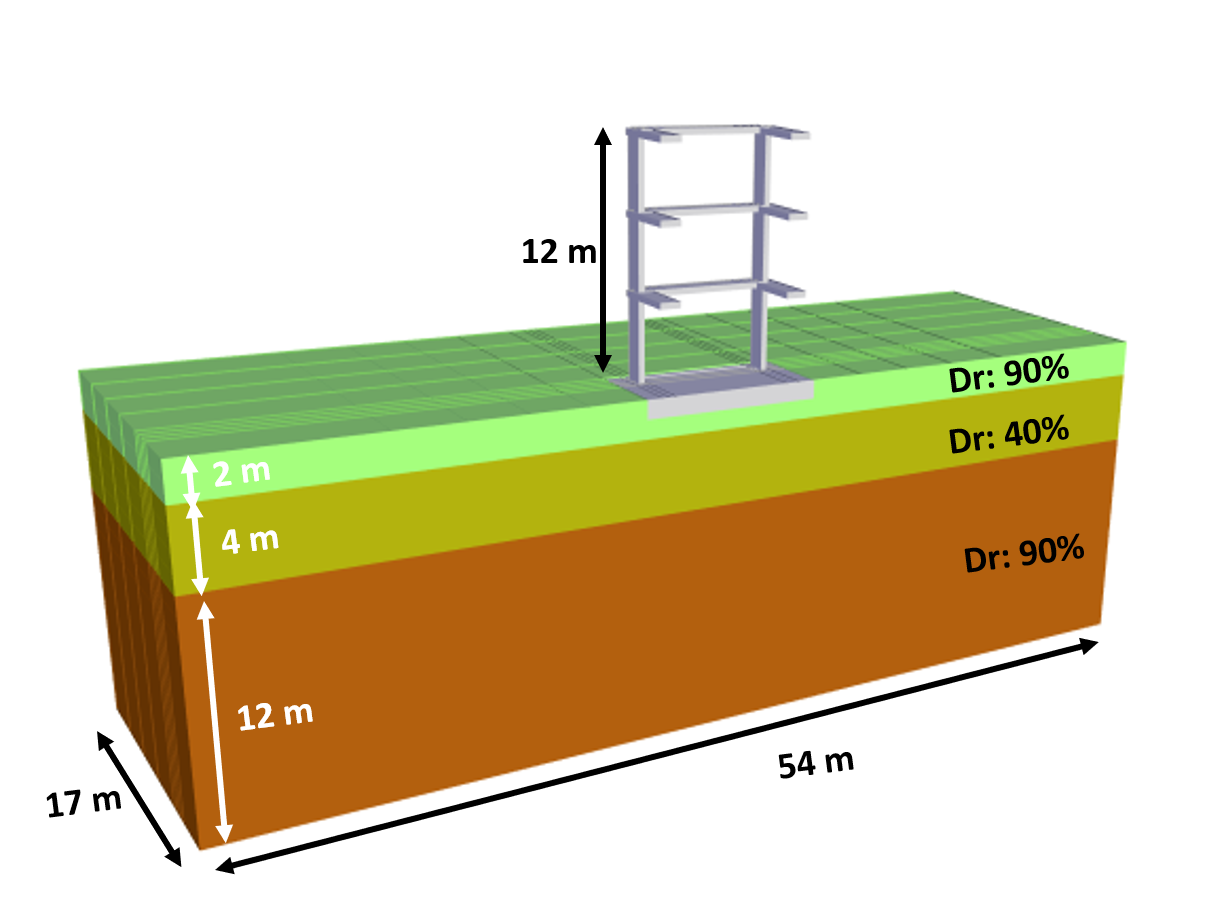
The use case workflow involves the following steps:
-
Creating the OpenSees input files using STKO.
-
Submitting the OpenSees job to the Stampede2 HPC resources at DesignSafe/TACC.
-
Post-processing the results using STKO and python.
Create OpenSees Model using STKO
- The user can create the input OpenSees-STKO model (both the 'main.tcl' and 'analysis_steps.tcl' files, as well as the '*.cdata' files) using STKO , which is available from the Visualization tab of the Tools & Applications section of the DesignSafe Workspace.
- Save all the files (tcl script and mpco.cdata files) in a folder under the user's My Data directory within the Data Depot.
- Alternatively, the input OpenSees-STKO model can be created on the user's local computer and all the files uploaded to a My Data folder.

Setup and submit OpenSees job via Jupyter notebook
A Jupyter notebook, SSI_MainDriver.ipynb , is provided that submits a job to the STKO compatible version of OpenSeesMP.
This Jupyter notebook utilizes the input file 'main.tcl', as well as 'analysis_steps.tcl' and the associated '*.cdata' files created by STKO. All of these files must be located in the same folder within the My Data directory of the DesignSafe Data Depot.
Setup job description
This script demonstrates how to use the agavepy SDK that uses the TAPIS API to setup the job description for the OpenSeesMP (V 3.0) App that is integrated with STKO. More details of using TAPIS API for enabling workflows in Jupyter notebook can be found in the DesignSafe webinar: Leveraging DesignSafe with TAPIS
- The user should edit the "job info" parameters as needed.
- The "control_jobname" should be modified to be meaningful for your analysis.
- The "control_processorsnumber" should be equal to the "number of partitions" in the STKO model.
from agavepy.agave import Agave
ag = Agave.restore()
import os
### Running OPENSEESMP (V 3.0)-STKO ver. 3.0.0.6709
app_name = 'OpenSeesMP'
app_id = 'opensees-mp-stko-3.0.0.6709u1'
storage_id = 'designsafe.storage.default'
##### One can revise the following job info ####
control_batchQueue = 'normal'
control_jobname = 'SSI_NM_Northridge_0913'
control_nodenumber = '1'
control_processorsnumber = '36'
control_memorypernode = '1'
control_maxRunTime = '24:00:00'Submit and Run job on DesignSafe
The script below submits the job to the HPC system.
job = ag.jobs.submit(body=job_description)
print("Job launched")import time
status = ag.jobs.getStatus(jobId=job["id"])["status"]
while status != "FINISHED":
status = ag.jobs.getStatus(jobId=job["id"])["status"]
print(f"Status: {status}")
time.sleep(3600)Identify Job ID and Archived Location
After completing the analysis, the results are saved to an archive directory. This script fetches the jobID and identifies the path of the archived location on DS.
jobinfo = ag.jobs.get(jobId=job.id)
jobinfo.archivePath
user = jobinfo.archivePath.split('/', 1)[0]jobinfo.archivePath
'$username/archive/jobs/job-3511e755-cb3f-4e3e-92b9-615cc40d39e6-007'Post-processing on DesignSafe
The output from an OpenSeesMP-STKO analysis are provided in a number of '*.mpco' files, and these files can be visualized and data values extracted using STKO. Additionally, the user can manually add recorders to the 'analysis_steps.tcl' file created by STKO and the output from these recorders will be saved in *.txt files. These *.txt can be imported into Jupyter for post-processing, visualization, and plotting.
Visualize and extract data from STKO
After the job is finished, the user can use STKO to visualize the results in the '*.mpco' files that are located in the archive directory. If the user would like to extract data from the GUI of STKO, they can copy and paste the data using the "Leafpad" text editor within the DS virtual machine that serves STKO. The user can then save the text file to a folder within the user's My Data directory.
Example post-processing scripts using Jupyter
A separate Jupyter notebook is provided (Example post-processing scripts.ipynb ) that post-processes data from OpenSees recorders and save in *.txt files. The Jupyter notebook is set up to open the *.txt files after thay have been copied from the archive directory to the same My Data in which the notebook resides.
For this example, recorders are created to generate output presented in terms of:
-
Acceleration response spectra at the mid of loose sand layer, foundation, and roof.
-
Evolution of excess pore water pressure at the bottom, mid, and top of the loose sand layer.
-
Time history of foundation settlement and tilt.
Creating recorders
To manually add the recorders, the user needs to first identify the id of the nodes via STKO (see StruList and SoilList below) and their corresponding partition id (i.e., Process_id). These recorders should be added into the "analysis_steps.tcl" before running the model. Note that the "analysis_steps.tcl" is automatically generated by STKO.
if {$process_id == 20} {
set StruList {471 478 481}
eval "recorder Node -file $filename.txt -time -node $StruList -dof 1 3 -dT $timestep disp"
eval "recorder Node -file $filename.txt -time -node $StruList -dof 1 -dT $timestep accel"
set SoilList {2781}
eval "recorder Node -file $filename.txt -time -node $SoilList -dof 1 3 -dT $timestep disp"
eval "recorder Node -file $filename.txt -time -node $SoilList -dof 1 -dT $timestep accel"
eval "recorder Node -file $filename.txt -time -node $SoilList -dof 4 -dT $timestep vel"
}Example Post-processing Results
This section shows the results from the post-processing scripts performed via the Jupyter notebook. The notebook is broken into segments with explanations of each section of code. Users should edit the code to fit their own needs.
Response spectra for motions at various locations within the model
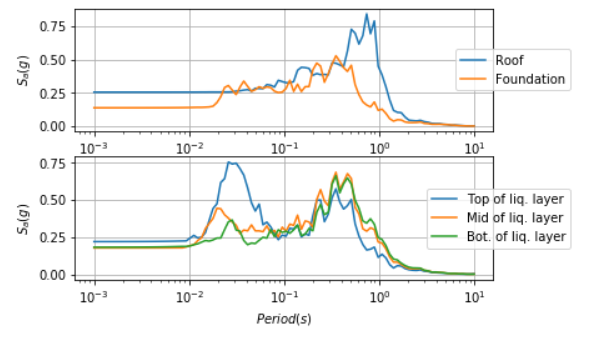
Time history of excess pore pressure at different locations in the soil
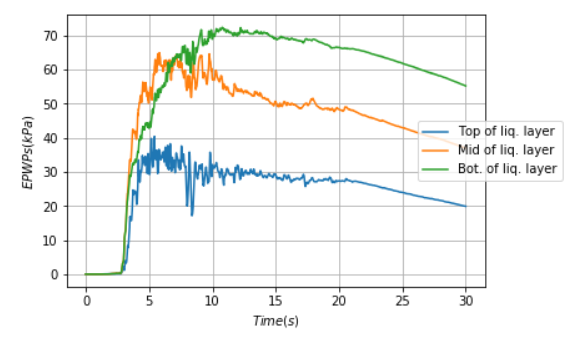
Time history of foundation settlement and tilt
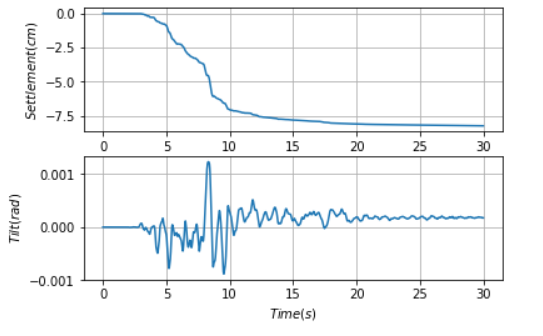
Experimental Shake Table Testing
Experimental Shake Table Testing
Integrated Workflow of Experiments using Jupyter Notebooks: From Experimental Design to Publication
Enrique Simbort - University of California, San Diego
Gilberto Mosqueda - University of California, San Diego
Key Words: Jupyter, Interactive Data Analysis, UCSD NHERI Facility
Jupyter Notebooks can provide fully integrated workflows of experiments from documentation of experimental design through analysis and publishing of data using the DesignSafe cyberinfrastructure. A series of Notebooks are being developed to demonstrate their use in the experimental workflow including notebooks showing how to view and analyzed past published data and data from testing of a reconfigurable, modular test bed building planned to be tested on the NHERI@UC San Diego Experimental Facility. The Python-based code is implemented in a modular fashion so that components can be used as desired in other experiments and are transferable to other experimental facilities. In the examples provided, the Notebook can be used to evaluate shake table performance as well as dynamic properties of the structure. A key functionality is to increase the integration and collaboration between researchers at local or remote sites to view and analyze the experimental data during and after testing including after the data is published. As Notebooks are developed to view experimental data by the research team, they can also be published with the data allowing other researchers to quickly view the data for promoting data reuse. Examples are providing for viewing data from past shake table experiments including NEES and more current NHERI data repositories.
Resources
Jupyter Notebooks
The following Jupyter notebooks are available to facilitate the analysis of each case. They are described in details in this section. You can access and run them directly on DesignSafe by clicking on the "Open in DesignSafe" button.
DesignSafe Resources
The following DesignSafe resources were used in developing this use case.
Background
Citation and Licensing
- Please cite Mosqueda et al. (2017) to acknowledge the use of resources from this use case with additional data sources referenced below.
- Please cite Rathje et al. (2017) to acknowledge the use of DesignSafe resources.
- This software is distributed under the GNU General Public License .
Description
As the cyberinfrastructure for The Natural Hazard Engineering Research Infrastructure (NHERI), DesignSafe, see Rathje et al. (2020) , provides a collaborative workspace for cloud-based data analysis, data sharing, curation and publication of models and data. Within this workspace, Jupyter Notebooks can be applied to perform data analysis in an interactive environment with access to published data. A rich set of data from natural hazard experiments and field studies is available from NHERI projects and its predecessor the Network for Earthquake Engineering Simulation (NEES). Since one of the major goals of DesignSafe is to provide a collaborative workspace by means of data sharing and access for data reuses, the main objective of this document is to demonstrate the use of Jupyter Notebook for viewing and analyzing published data using cloud-based tools.
This use case includes a series of Jupyter Notebooks aimed to serve as a learning tool for viewing and analyzing data from shake table experiments including:
-
The first module examines the performance of Hybrid Simulation Experiments conducted on the 1D Large High Performance Outdoor Shake table at UC San Diego with the data published in DesignSafe by Vega et al. (2018) . This module focuses on the response of the shake table including tools to compare different signals. Data extraction and processing of measured sensor data includes comparison of time history signals, comparison of signals in the frequency domain using FFT and comparison of response spectra that show for example target and measured table response.
-
The second module examines the use of Jupyter Notebooks including Python libraries for structural response and system identification. In this case data from a past NEES experiment conducted by Mosqueda et al. (2017) of three-story moment frame structure is examined. The data published in DataDepot under as a NEES project. Using selected sensors at each story of the structure and white noise excitation, the frequencies and mode shapes of the structure are identified. The processing tools rely on existing libraries in Python demonstrating the wealth of access to subroutine that can be applied for analysis.
-
The third module is a Jupyter notebook for viewing and analyzing data from tests on a 3-story steel Modular Testbed Building (MTB2) conducted on the recently upgrades 6DOF shake table at the UC San Diego NHEIR Experimental Facilty as reported by Van Den Einde et al (2020) . These tests will examine the shake table performance and structural response for the 3D structure. These tests are currently in progress with the notebook under development concurrently.
Jupyter Notebooks for Experimental Data
Jupyter Notebooks work as an interactive development environment to code and view data in a report format. Within the notebook, the combination of cells enables formatted text and interactive plotting for viewing and analyzing data. Users can select data files and data channels for viewing and processing with the ability to view and print complete reports. Jupyter Notebooks are accessible in DesignSafe through the workspace analysis tools and can access private or public data in Data Depot. Sample modules are presented here that have been developed using published data in Data Depot including those by Vega et al. (2018) and Masroor et al. (2010) . These modules will be configured and applied within the workflow of the MTB2 during shakedown testing.
Case 1. Shake table performance
A set of modules have been developed to evaluate the performance of the shake table using data from past experiments conducted to demonstrate the hybrid testing capabilities of LHPOST, see Vega et al (2020) . For these hybrid tests, separate Jupyter Notebooks have been developed to consider the various sources of generated data including i) Shake Table Controller, ii) the primary Data Acquisition System (DAQ), and iii) additional computational sources for hybrid testing. In a typical shake table test, the first two sources of data would be included plus any other user specified data acquisition system.
Data collected by the shake table controller is expected to be standard across most tests and useful to verify the performance of the shake table in reproducing the ground motions. Here, data from the shake table controller is used to compare reference command and measured feedback data to evaluate the fidelity of the shake table in reproducing the desired ground motions, see Vega et al. (2018) for an example. The Jupyter notebook functionality includes interactive plotters for viewing either a single channel or multiple channels to compare the reference input and feedback, for example, by viewing the time history, Fourier Transform or Response Spectra (Fig 1). The shake table controller sampling rate was set to a frequency of 2048 [Hz] for this test. Initial implementation of the code required about 3.5 minutes to run. To improve the run-time, various options were explored including down sampling and use of tools such as those being developed by Brandenberg, S., J., & Yang, Y. (2021) to calculate the spectral acceleration. By using these tools, the run time was reduced to approximately 10 s. The module was implemented for the previous 1-D capability of LHPOST but can be easily extended for its newly upgraded 6DOF capabilities.

Figure 1. Evaluation of shake table performance through comparison of command reference and feedback including a) Fourier Transform and b) response spectra .
Case 2. Module for Structural Response and System Identification
The primary goal of the structural response module is to quickly and accurately analyze experimental data. For development and testing of available algorithms, experimental data from a previous dynamic experiment involving a ¼ scale three-story steel moment frame structure were used. For detailed information see Mosqueda et al. (2017) , with the data available in Masroor et al. (2010) . A cross spectral density function (CSD) is applied to compare the white noise acceleration input at the platen to the acceleration at each floor. To improve code clarity and compatibility for future investigators, the CSD function from the SciPy signal package, developed by Virtanen et al (2020) , is implemented, which is well documented. The CSD for each floor is plotted using the matplotlib library. The resulting CSD plots are shown up to 20 Hz in Fig. 2 and identify the natural frequencies of the structure. Modal displacements can also be calculated directly from the CSD function outputs. This is accomplished by using the frequency-power relation between acceleration spectral density functions and displacement spectral density functions. The modal displacements for each story occur at frequencies where the CSD has a local maximum. To obtain these values for the test data of the three-story structure, the frequencies of the first three local maxima were recorded. For future use of this code, the desired number of mode shapes can be scaled by adding or removing local maxima terms at the start of the mode shapes code section. Using the CSD function does not take into account the sign of the modal displacement, however, since these functions are strictly positive over their domain. To account for this, the output of the CSD function at the local maxima frequencies is reexamined without considering the absolute values of its components to identify if the parameters yield a negative number at the corresponding frequency. The rough shape of the modal displacements is plotted as shown in Fig. 3. Future work for this notebook includes generating a smoothing function for the mode shapes and comparison of data from different tests to identify changes in dynamic properties through the testing series that could be indicative of damage.
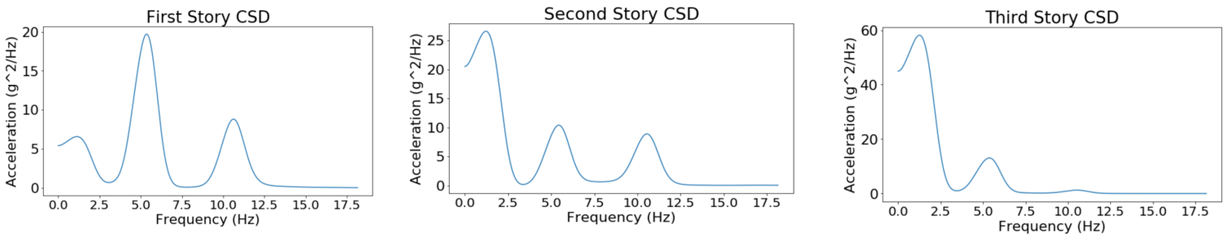
Figure 2. System identification of three story moment frame, by Masroor et al. (2010) , subjected to white noise from CSD function outputs.

Figure 3. Mode shapes estimation from 3-story building subjected to white noise on shake table.
Case 3. Integration of Notebooks in Experimental Workflow
The primary goal of this module is to develop the Jupyter Notebooks through the experimental program. The experiments are in planning and thus, this module would be programmed based on draft instrumentation plans. This module will plot the primary structural response such as story accelerations and drifts as well as employ system identification routines available in Python and previously demonstrated. Current work is exploring use of machine learning libraries for applications to these modules.
The Modular Testbed Building (MTB2), described in Morano et al. (2021), is designed to be a shared-use, reconfigurable experimental structure. The standard 3-story building can simulate braced frame and moment frame behavior through replaceable fuse type components including buckling restrained braced frames and Durafuse shear plate connections, respectively. The unique connection scheme allows for yielded fuse type members to be easily replaced to restore the structure to its original condition. The MTB2 can be constructed in various configurations with three examples shown in Fig 4. The lateral framing system in the 2-bay direction can be modeled as moment frames or braced frames. The single bay direction has a span of 20 feet and is a braced frame. Each span in the double bay direction is 16 feet. The story height for all floors is 12 feet with columns that extend 4 feet above the top floor. The Special Moment Frame (SMF) configuration utilizes replaceable shear fuse plates while the braced frame utilizes Buckling-Restrained Braces.

Figure 4. MTB2 building: a) SMF configuration (left), b) BRB-1 configuration (center), and c) BRB-2 configuration (right).
Summary
The Jupyter Notebooks developed for use through DesignSafe will facilitate the viewing and analysis of data sharing with collaborators from testing through data publication. A key advantage is the cloud-based approach that facilitates interactive data viewing and analysis in a report format without having to download large datasets. These tools are intended to make data more readily accessible and promote data reuse. The Jupyter Notebook presented here includes routines to evaluate the performance of the shake table and carry out system identification of structural models.
References
- Rathje et al. (2020) “Enhancing research in natural hazard engineering through the DesignSafe cyberinfrastructure”. Frontiers in Built Environment, 6:213.
- Vega et al (2020) “Implementation of real-time hybrid shake table testing using the UCSD large high-performance outdoor shake table”. Int. J. Lifecycle Performance Engineering, Vol. 4, p.80-102.
- Vega et al. (2018) "Five story building with tunned mass damper", in NHERI UCSD Hybrid Simulation Commissioning. DesignSafe-CI.
- Mosqueda et al. (2017) “Seismic Response of Base Isolated Buildings Considering Pounding to Moat Walls”. Technical report MCEER-13-0003.
- Van Den Einde et al (2020) “NHERI@ UC San Diego 6-DOF Large High-Performance Outdoor Shake Table Facility.” Frontiers in Built Environment, 6:181.
- Masroor et al. (2010) "Limit State Behavior of Base Isolated Structures: Fixed Base Moment Frame", DesignSafe-CI.
- Vega et al (2020), “Implementation of real-time hybrid shake table testing using the UCSD large high-performance outdoor shake table”. Int. J. Lifecycle Performance Engineering, Vol. 4, p.80-102.
- Brandenberg, S., J., & Yang, Y. (2021) "ucla_geotech_tools: A set of Python packages developed by the UCLA geotechnical group" (Version 1.0.2) [Computer software].
- Virtanen et al (2020) “SciPy 1.0: Fundamental Algorithms for Scientific Computing in Python”. Nature Methods, 17, 261–272.
- Chopra A. K., Dynamics of Structures. Harlow: Pearson Education, 2012.
- Morano M., Liu J., Hutchinson T. C., and Pantelides C.P. (2021), “Design and Analysis of a Modular Test Building for the 6-DOF Large High-Performace Outdoor Shake Table”, 17th World Conference on Earthquake Engineering, Japan.
- Mosqueda G, Guerrero N, Schmemmer Z., Lin L, Morano M., Liu J., Hutchinson T, Pantelides C P. Jupyter Notebooks for Data workflow of NHERI Experimental Facilities. Proceedings of the 12th National Conference in Earthquake Engineering, Earthquake Engineering Research Institute, Salt Lake City, UT. 2022.

